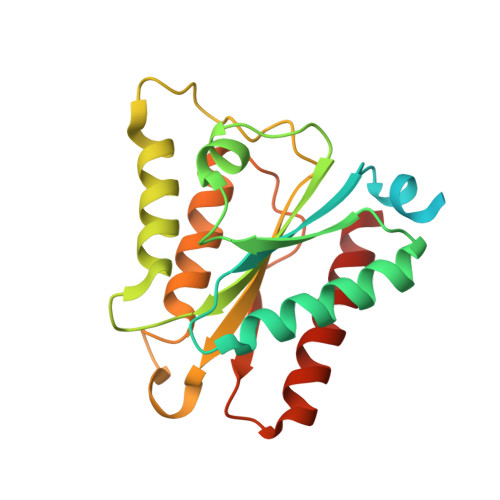
Biochemical and structural characterization of the flavodoxin-like domain of the Schizosaccharomyces japonicus putative tRNAPhe 4-demethylwyosine synthase Tyw1 in complex with FMN.
Sjekloca, L., Ferre-D'Amare, A.R.(2022) MicroPubl Biol 2022
- PubMed: 35693892 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.17912/micropub.biology.000570
- Primary Citation Related Structures:
6PUP, 6PUQ - PubMed Abstract:
The S-adenosyl-L-methionine-dependent tRNA 4-demethylwyosine synthase TYW1 catalyzes biosynthesis of 4-demethylwyosine (imG-14), the precursor for wyosine, the hypermodified guanine-derived nucleotide present at position 37 of phenylalanine tRNAs of archaea and eukarya. Eukaryotic TYW1 enzymes contain N-terminal flavodoxin-like and C-terminal radical-SAM domains. We determined co-crystal structures of the flavodoxin-like domain of the putative Tyw1 from Schizosaccharomyces japonicus in complex with flavin mononucleotide (FMN), exploiting an unexpected anomalous scatterer present in the recombinant protein. Our results show how eukaryotic TYW1 enzymes bind the coenzyme FMN and will help further elucidation of the structural enzymology of 4-demethylwyosine synthesis.
- Biochemistry and Biophysics Center, National Heart, Lung and Blood Institute, 50 South Drive, Bethesda, Maryland, 20892-8012, United States.
Organizational Affiliation: