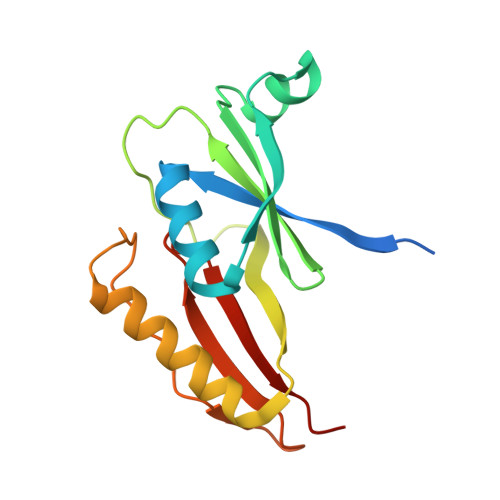
Crystal Structures of L-DOPA Dioxygenase fromStreptomyces sclerotialus.
Wang, Y., Shin, I., Fu, Y., Colabroy, K.L., Liu, A.(2019) Biochemistry 58: 5339-5350
- PubMed: 31180203 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acs.biochem.9b00396
- Primary Citation Related Structures:
6ON1, 6ON3 - PubMed Abstract:
Extradiol dioxygenases are essential biocatalysts for breaking down catechols. The vicinal oxygen chelate (VOC) superfamily contains a large number of extradiol dioxygenases, most of which are found as part of catabolic pathways degrading a variety of natural and human-made aromatic rings. The l-3,4-dihydroxyphenylalanine (L-DOPA) extradiol dioxygenases compose a multitude of pathways that produce various antibacterial or antitumor natural products. The structural features of these dioxygenases are anticipated to be distinct from those of other VOC extradiol dioxygenases. Herein, we identified a new L-DOPA dioxygenase from the thermophilic bacterium Streptomyces sclerotialus (SsDDO) through a sequence and genome context analysis. The activity of SsDDO was kinetically characterized with L-DOPA using an ultraviolet-visible spectrophotometer and an oxygen electrode. The optimal temperature of the assay was 55 °C, at which the K m and k cat of SsDDO were 110 ± 10 μM and 2.0 ± 0.1 s -1 , respectively. We determined the de novo crystal structures of SsDDO in the ligand-free form and as a substrate-bound complex, refined to 1.99 and 2.31 Å resolution, respectively. These structures reveal that SsDDO possesses a form IV arrangement of βαβββ modules, the first characterization of this assembly from among the VOC/type I extradiol dioxygenase protein family. Electron paramagnetic resonance spectra of Fe-NO adducts for the resting and substrate-bound enzyme were obtained. This work contributes to our understanding of a growing class of topologically distinct VOC dioxygenases, and the obtained structural features will improve our understanding of the extradiol cleavage reaction within the VOC superfamily.
- Department of Chemistry , University of Texas at San Antonio , San Antonio , Texas 78249 , United States.
Organizational Affiliation: