The Molecular Control of Calcitonin Receptor Signaling.
Dal Maso, E., Glukhova, A., Zhu, Y., Garcia-Nafria, J., Tate, C.G., Atanasio, S., Reynolds, C.A., Ramirez-Aportela, E., Carazo, J.M., Hick, C.A., Furness, S.G.B., Hay, D.L., Liang, Y.L., Miller, L.J., Christopoulos, A., Wang, M.W., Wootten, D., Sexton, P.M.(2019) Acs Pharmacol Transl Sci 2: 31-51
- PubMed: 32219215 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acsptsci.8b00056
- Primary Citation Related Structures:
6NIY - PubMed Abstract:
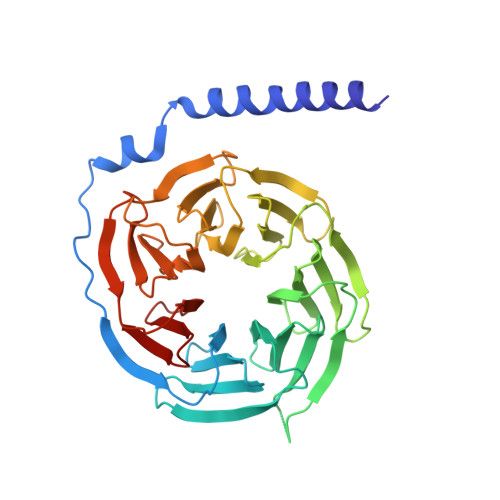
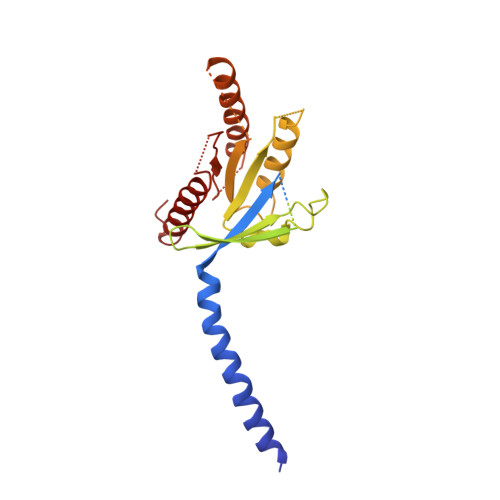
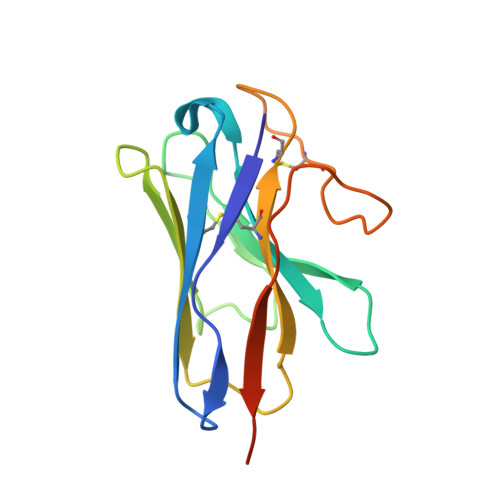
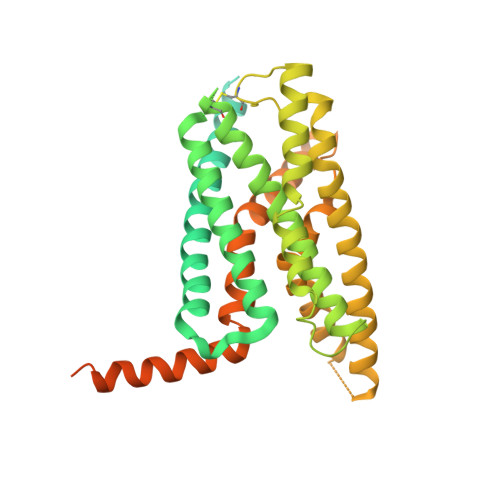
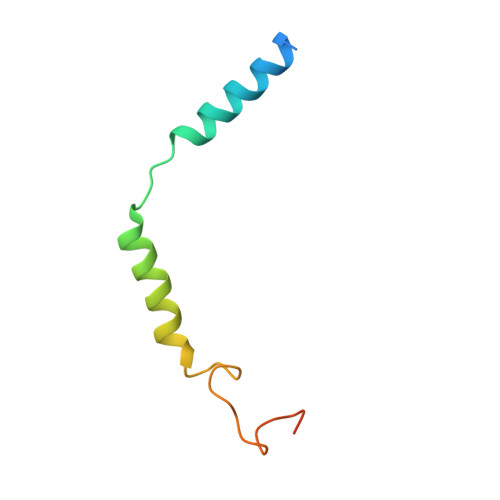
The calcitonin receptor (CTR) is a class B G protein-coupled receptor (GPCR) that responds to the peptide hormone calcitonin (CT). CTs are clinically approved for the treatment of bone diseases. We previously reported a 4.1 Å structure of the activated CTR bound to salmon CT (sCT) and heterotrimeric Gs protein by cryo-electron microscopy (Liang, Y.-L., et al . Phase-plate cryo- EM structure of a class B GPCR-G protein complex. Nature 2017 , 546 , 118-123). In the current study, we have reprocessed the electron micrographs to yield a 3.3 Å map of the complex. This has allowed us to model extracellular loops (ECLs) 2 and 3, and the peptide N-terminus that previously could not be resolved. We have also performed alanine scanning mutagenesis of ECL1 and the upper segment of transmembrane helix 1 (TM1) and its extension into the receptor extracellular domain (TM1 stalk), with effects on peptide binding and function assessed by cAMP accumulation and ERK1/2 phosphorylation. These data were combined with previously published alanine scanning mutagenesis of ECL2 and ECL3 and the new structural information to provide a comprehensive 3D map of the molecular surface of the CTR that controls binding and signaling of distinct CT and related peptides. The work highlights distinctions in how different, related, class B receptors may be activated. The new mutational data on the TM1 stalk and ECL1 have also provided critical insights into the divergent control of cAMP versus pERK signaling and, collectively with previous mutagenesis data, offer evidence that the conformations linked to these different signaling pathways are, in many ways, mutually exclusive. This study furthers our understanding of the complex nature of signaling elicited by GPCRs and, in particular, that of the therapeutically important class B subfamily.
- Drug Discovery Biology, Monash Institute of Pharmaceutical Sciences, Monash University, Parkville 3052, Victoria Australia.
Organizational Affiliation: