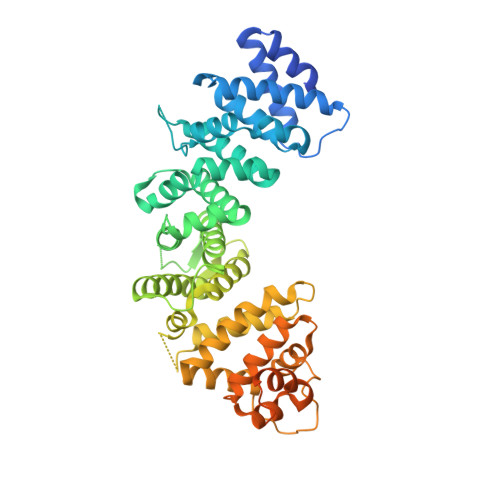
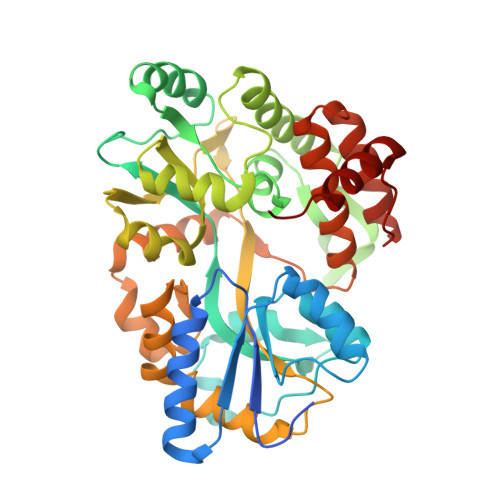
Structural underpinnings of Ric8A function as a G-protein alpha-subunit chaperone and guanine-nucleotide exchange factor.
Srivastava, D., Gakhar, L., Artemyev, N.O.(2019) Nat Commun 10: 3084-3084
- PubMed: 31300652 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-019-11088-x
- Primary Citation Related Structures:
6N84, 6N85, 6N86 - PubMed Abstract:
Resistance to inhibitors of cholinesterase 8A (Ric8A) is an essential regulator of G protein α-subunits (Gα), acting as a guanine nucleotide exchange factor and a chaperone. We report two crystal structures of Ric8A, one in the apo form and the other in complex with a tagged C-terminal fragment of Gα. These structures reveal two principal domains of Ric8A: an armadillo-fold core and a flexible C-terminal tail. Additionally, they show that the Gα C-terminus binds to a highly-conserved patch on the concave surface of the Ric8A armadillo-domain, with selectivity determinants residing in the Gα sequence. Biochemical analysis shows that the Ric8A C-terminal tail is critical for its stability and function. A model of the Ric8A/Gα complex derived from crosslinking mass spectrometry and molecular dynamics simulations suggests that the Ric8A C-terminal tail helps organize the GTP-binding site of Gα. This study lays the groundwork for understanding Ric8A function at the molecular level.
- Department of Molecular Physiology and Biophysics, University of Iowa Carver College of Medicine, Iowa City, IA, 52242, USA.
Organizational Affiliation: