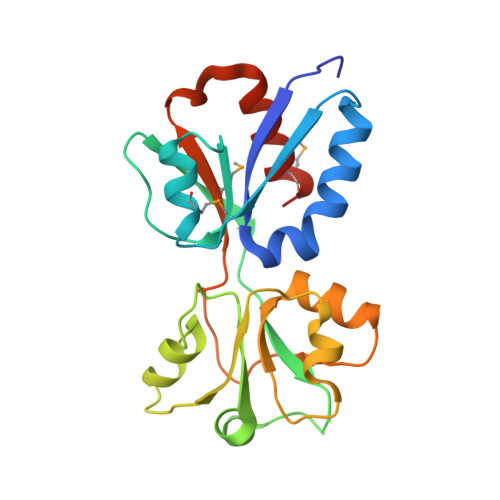
Crystal Structure of the Regulatory Domain of MexT, a Transcriptional Activator of the MexEFOprN Efflux Pump inPseudomonas aeruginosa.
Kim, S., Kim, S.H., Ahn, J., Jo, I., Lee, Z.W., Choi, S.H., Ha, N.(2019) Mol Cells 42: 850-857
- PubMed: 31722511 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.14348/molcells.2019.0168
- Primary Citation Related Structures:
6L33 - PubMed Abstract:
The Gram-negative opportunistic pathogen, Pseudomonas aeruginosa , has multiple multidrug efflux pumps. MexT, a LysR-type transcriptional regulator, functions as a transcriptional activator of the MexEF-OprN efflux system. MexT consists of an N-terminal DNA-binding domain and a C-terminal regulatory domain (RD). Little is known regarding MexT ligands and its mechanism of activation. We elucidated the crystal structure of the MexT RD at 2.0 Å resolution. The structure comprised two protomer chains in a dimeric arrangement. MexT possessed an arginine-rich region and a hydrophobic patch lined by a variable loop, both of which are putative ligand-binding sites. The three-dimensional structure of MexT provided clues to the interacting ligand structure. A DNase I footprinting assay of full-length MexT identified two MexT-binding sequence in the mexEF oprN promoter. Our findings enhance the understanding of the regulation of MexT-dependent activation of efflux pumps.
- Department of Agricultural Biotechnology, Center for Food Safety and Toxicology, Center for Food and Bioconvergence, and Research Institute for Agriculture and Life Sciences, Seoul National University, Seoul 08826, Korea.
Organizational Affiliation: