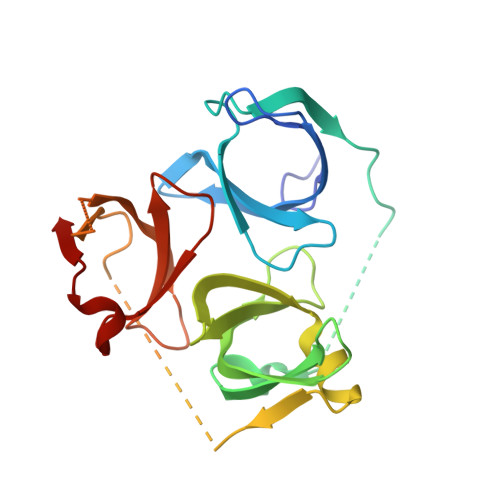
A Chemical Probe for Tudor Domain Protein Spindlin1 to Investigate Chromatin Function.
Fagan, V., Johansson, C., Gileadi, C., Monteiro, O., Dunford, J.E., Nibhani, R., Philpott, M., Malzahn, J., Wells, G., Faram, R., Cribbs, A.P., Halidi, N., Li, F., Chau, I., Greschik, H., Velupillai, S., Allali-Hassani, A., Bennett, J., Christott, T., Giroud, C., Lewis, A.M., Huber, K.V.M., Athanasou, N., Bountra, C., Jung, M., Schule, R., Vedadi, M., Arrowsmith, C., Xiong, Y., Jin, J., Fedorov, O., Farnie, G., Brennan, P.E., Oppermann, U.(2019) J Med Chem 62: 9008-9025
- PubMed: 31550156 Search on PubMed
- DOI: https://doi.org/10.1021/acs.jmedchem.9b00562
- Primary Citation Related Structures:
6I8B, 6I8L, 6I8Y - PubMed Abstract:
Modifications of histone tails, including lysine/arginine methylation, provide the basis of a "chromatin or histone code". Proteins that contain "reader" domains can bind to these modifications and form specific effector complexes, which ultimately mediate chromatin function. The spindlin1 (SPIN1) protein contains three Tudor methyllysine/arginine reader domains and was identified as a putative oncogene and transcriptional coactivator. Here we report a SPIN1 chemical probe inhibitor with low nanomolar in vitro activity, exquisite selectivity on a panel of methyl reader and writer proteins, and with submicromolar cellular activity. X-ray crystallography showed that this Tudor domain chemical probe simultaneously engages Tudor domains 1 and 2 via a bidentate binding mode. Small molecule inhibition and siRNA knockdown of SPIN1, as well as chemoproteomic studies, identified genes which are transcriptionally regulated by SPIN1 in squamous cell carcinoma and suggest that SPIN1 may have a role in cancer related inflammation and/or cancer metastasis.
- Structural Genomics Consortium, Nuffield Department of Medicine , University of Oxford , OX3 7DQ Oxford , U.K.
Organizational Affiliation: