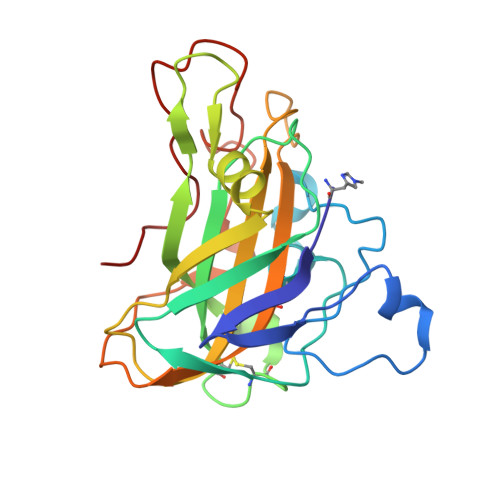
Structure of a lytic polysaccharide monooxygenase from Aspergillus fumigatus and an engineered thermostable variant.
Lo Leggio, L., Weihe, C.D., Poulsen, J.N., Sweeney, M., Rasmussen, F., Lin, J., De Maria, L., Wogulis, M.(2018) Carbohydr Res 469: 55-59
- PubMed: 30296642 Search on PubMed
- DOI: https://doi.org/10.1016/j.carres.2018.08.009
- Primary Citation Related Structures:
6H1Z, 6HA5, 6HAQ - PubMed Abstract:
Lytic polysaccharide monooxygenases (LPMOs) are industrial enzymes which are gaining use in second generation bioethanol production from lignocellulose by acting in synergy with glycoside hydrolases. Here we present the X-ray crystal structure of an AA9 fungal LPMO from Aspergillus fumigatus and a variant which has been shown to have better performance at elevated temperatures. Based on the structures, thermal denaturation data and theoretical calculations, we provide a suggestion for the structural basis of the improved stability.
- Department of Chemistry, University of Copenhagen, Universitetsparken 5, 2100, Copenhagen, Denmark. Electronic address: leila@chem.ku.dk.
Organizational Affiliation: