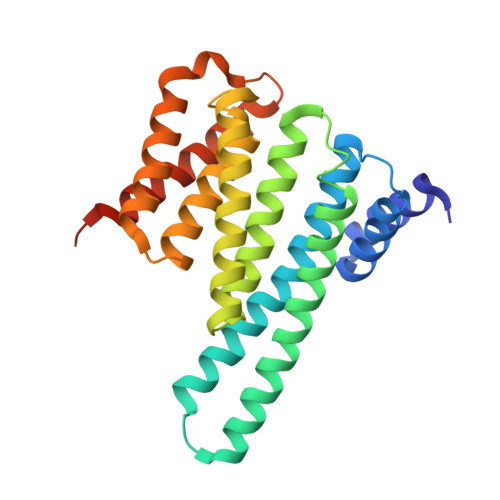
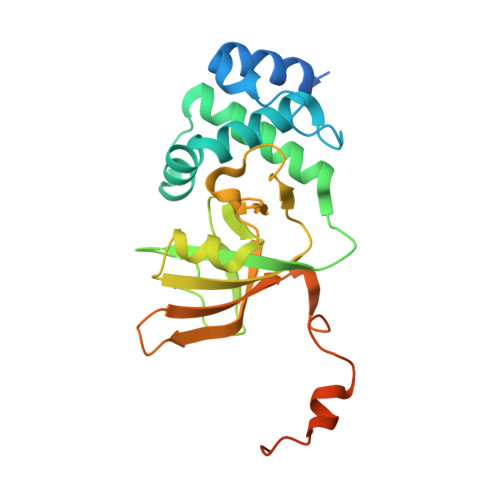
14-3-3 proteins activate Pseudomonas exotoxins-S and -T by chaperoning a hydrophobic surface.
Karlberg, T., Hornyak, P., Pinto, A.F., Milanova, S., Ebrahimi, M., Lindberg, M., Pullen, N., Nordstrom, A., Loverli, E., Caraballo, R., Wong, E.V., Nareoja, K., Thorsell, A.G., Elofsson, M., De La Cruz, E.M., Bjorkegren, C., Schuler, H.(2018) Nat Commun 9: 3785-3785
- PubMed: 30224724 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-018-06194-1
- Primary Citation Related Structures:
6GN0, 6GN8, 6GNJ, 6GNK, 6GNN - PubMed Abstract:
Pseudomonas are a common cause of hospital-acquired infections that may be lethal. ADP-ribosyltransferase activities of Pseudomonas exotoxin-S and -T depend on 14-3-3 proteins inside the host cell. By binding in the 14-3-3 phosphopeptide binding groove, an amphipathic C-terminal helix of ExoS and ExoT has been thought to be crucial for their activation. However, crystal structures of the 14-3-3β:ExoS and -ExoT complexes presented here reveal an extensive hydrophobic interface that is sufficient for complex formation and toxin activation. We show that C-terminally truncated ExoS ADP-ribosyltransferase domain lacking the amphipathic binding motif is active when co-expressed with 14-3-3. Moreover, swapping the amphipathic C-terminus with a fragment from Vibrio Vis toxin creates a 14-3-3 independent toxin that ADP-ribosylates known ExoS targets. Finally, we show that 14-3-3 stabilizes ExoS against thermal aggregation. Together, this indicates that 14-3-3 proteins activate exotoxin ADP-ribosyltransferase domains by chaperoning their hydrophobic surfaces independently of the amphipathic C-terminal segment.
- Department of Biosciences and Nutrition, Karolinska Institutet, Hälsovägen 4c, 14157, Huddinge, Sweden.
Organizational Affiliation: