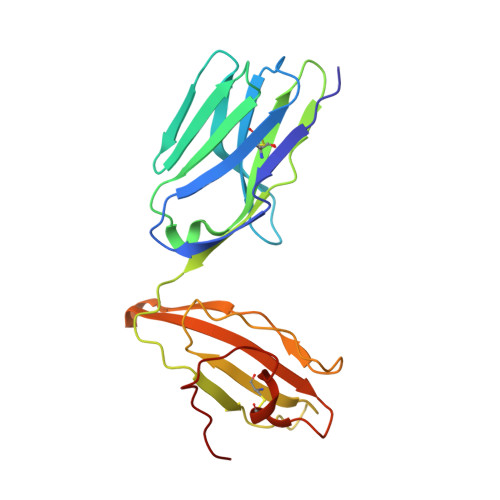
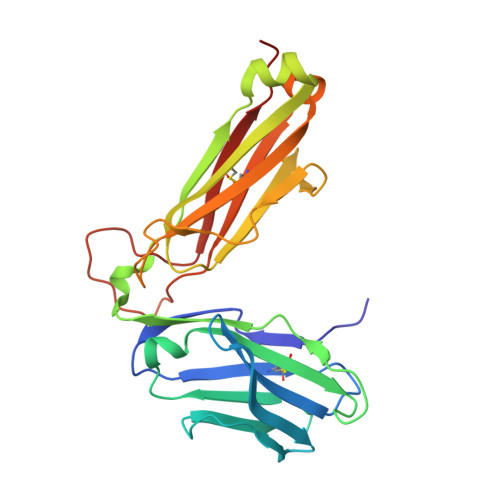
In Silicoand Structural Analyses Demonstrate That Intrinsic Protein Motions Guide T Cell Receptor Complementarity Determining Region Loop Flexibility.
Holland, C.J., MacLachlan, B.J., Bianchi, V., Hesketh, S.J., Morgan, R., Vickery, O., Bulek, A.M., Fuller, A., Godkin, A., Sewell, A.K., Rizkallah, P.J., Wells, S., Cole, D.K.(2018) Front Immunol 9: 674-674
- PubMed: 29696015 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.3389/fimmu.2018.00674
- Primary Citation Related Structures:
6EH4, 6EH5, 6EH6, 6EH7, 6EH8, 6EH9, 6FR3, 6FR4, 6FR5, 6FR6, 6FR7, 6FR8, 6FR9, 6FRA, 6FRB, 6FRC, 6FUM, 6FUN, 6FUO, 6FUP, 6FUQ, 6FUR - PubMed Abstract:
T-cell immunity is controlled by T cell receptor (TCR) binding to peptide major histocompatibility complexes (pMHCs). The nature of the interaction between these two proteins has been the subject of many investigations because of its central role in immunity against pathogens, cancer, in autoimmunity, and during organ transplant rejection. Crystal structures comparing unbound and pMHC-bound TCRs have revealed flexibility at the interaction interface, particularly from the perspective of the TCR. However, crystal structures represent only a snapshot of protein conformation that could be influenced through biologically irrelevant crystal lattice contacts and other factors. Here, we solved the structures of three unbound TCRs from multiple crystals. Superposition of identical TCR structures from different crystals revealed some conformation differences of up to 5 Å in individual complementarity determining region (CDR) loops that are similar to those that have previously been attributed to antigen engagement. We then used a combination of rigidity analysis and simulations of protein motion to reveal the theoretical potential of TCR CDR loop flexibility in unbound state. These simulations of protein motion support the notion that crystal structures may only offer an artifactual indication of TCR flexibility, influenced by crystallization conditions and crystal packing that is inconsistent with the theoretical potential of intrinsic TCR motions.
- Division of Infection and Immunity and Systems Immunity Research Institute, Cardiff University School of Medicine, Cardiff, United Kingdom.
Organizational Affiliation: