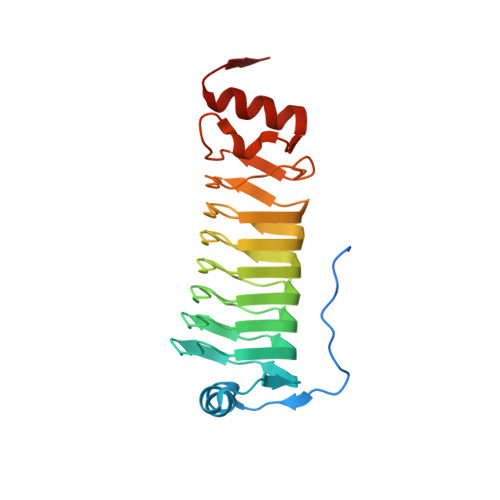
Cotranslational Folding of a Pentarepeat beta-Helix Protein.
Notari, L., Martinez-Carranza, M., Farias-Rico, J.A., Stenmark, P., von Heijne, G.(2018) J Mol Biology 430: 5196-5206
- PubMed: 30539762 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2018.10.016
- Primary Citation Related Structures:
6FLS - PubMed Abstract:
It is becoming increasingly clear that many proteins start to fold cotranslationally before the entire polypeptide chain has been synthesized on the ribosome. One class of proteins that a priori would seem particularly prone to cotranslational folding is repeat proteins, that is, proteins that are built from an array of nearly identical sequence repeats. However, while the folding of repeat proteins has been studied extensively in vitro with purified proteins, only a handful of studies have addressed the issue of cotranslational folding of repeat proteins. Here, we have determined the structure and studied the cotranslational folding of a β-helix pentarepeat protein from the human pathogen Clostridium botulinum-a homolog of the fluoroquinolone resistance protein MfpA-using an assay in which the SecM translational arrest peptide serves as a force sensor to detect folding events. We find that cotranslational folding of a segment corresponding to the first four of the eight β-helix coils in the protein produces enough force to release ribosome stalling and that folding starts when this unit is ~35 residues away from the P-site, near the distal end of the ribosome exit tunnel. An additional folding transition is seen when the whole PENT moiety emerges from the exit tunnel. The early cotranslational formation of a folded unit may be important to avoid misfolding events in vivo and may reflect the minimal size of a stable β-helix since it is structurally homologous to the smallest known β-helix protein, a four-coil protein that is stable in solution.
- Department of Biochemistry and Biophysics, Stockholm University, SE-106 91 Stockholm, Sweden.
Organizational Affiliation: