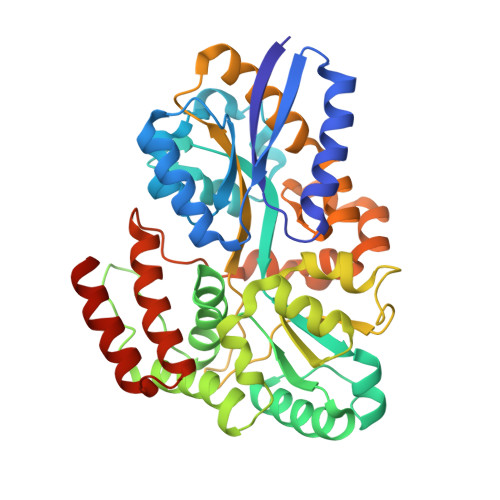
Differential Substrate Recognition by Maltose Binding Proteins Influenced by Structure and Dynamics.
Shukla, S., Bafna, K., Gullett, C., Myles, D.A.A., Agarwal, P.K., Cuneo, M.J.(2018) Biochemistry 57: 5864-5876
- PubMed: 30204415 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acs.biochem.8b00783
- Primary Citation Related Structures:
6DTQ, 6DTR, 6DTS, 6DTT, 6DTU - PubMed Abstract:
The genome of the hyperthermophile Thermotoga maritima contains three isoforms of maltose binding protein (MBP) that are high-affinity receptors for di-, tri-, and tetrasaccharides. Two of these proteins (tmMBP1 and tmMBP2) share significant sequence identity, approximately 90%, while the third (tmMBP3) shares less than 40% identity. MBP from Escherichia coli (ecMBP) shares 35% sequence identity with the tmMBPs. This subset of MBP isoforms offers an interesting opportunity to investigate the mechanisms underlying the evolution of substrate specificity and affinity profiles in a genome where redundant MBP genes are present. In this study, the X-ray crystal structures of tmMBP1, tmMBP2, and tmMBP3 are reported in the absence and presence of oligosaccharides. tmMBP1 and tmMBP2 have binding pockets that are larger than that of tmMBP3, enabling them to bind to larger substrates, while tmMBP1 and tmMBP2 also undergo substrate-induced hinge bending motions (∼52°) that are larger than that of tmMBP3 (∼35°). Small-angle X-ray scattering was used to compare protein behavior in solution, and computer simulations provided insights into dynamics of these proteins. Comparing quantitative protein-substrate interactions and dynamical properties of tmMBPs with those of the promiscuous ecMBP and disaccharide selective Thermococcus litoralis MBP provides insights into the features that enable selective binding. Collectively, the results provide insights into how the structure and dynamics of tmMBP homologues enable them to differentiate between a myriad of chemical entities while maintaining their common fold.
- Graduate School of Genome Science & Technology , The University of Tennessee , Knoxville , Tennessee 37996-0840 , United States.
Organizational Affiliation: