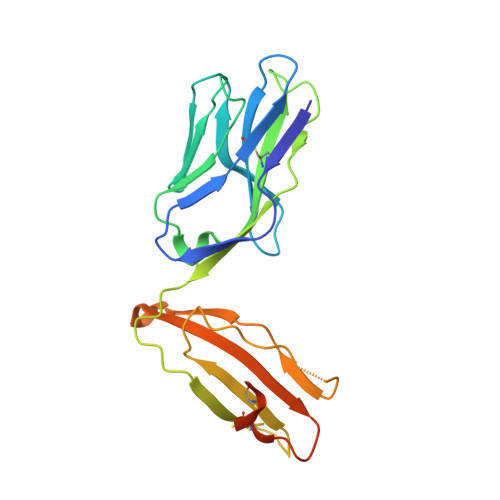
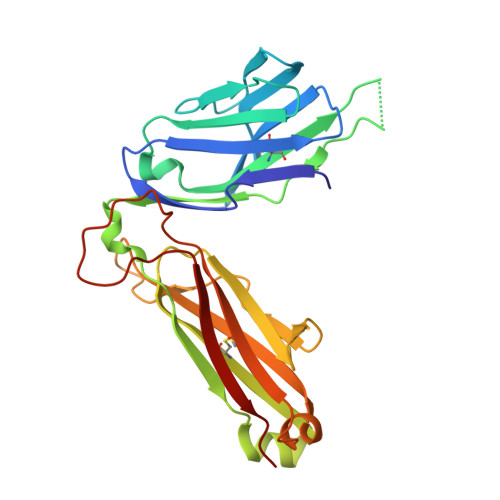
A T-cell receptor escape channel allows broad T-cell response to CD1b and membrane phospholipids.
Shahine, A., Reinink, P., Reijneveld, J.F., Gras, S., Holzheimer, M., Cheng, T.Y., Minnaard, A.J., Altman, J.D., Lenz, S., Prandi, J., Kubler-Kielb, J., Moody, D.B., Rossjohn, J., Van Rhijn, I.(2019) Nat Commun 10: 56-56
- PubMed: 30610190 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-018-07898-0
- Primary Citation Related Structures:
6CUG, 6CUH, 6D64 - PubMed Abstract:
CD1 proteins are expressed on dendritic cells, where they display lipid antigens to T-cell receptors (TCRs). Here we describe T-cell autoreactivity towards ubiquitous human membrane phospholipids presented by CD1b. These T-cells discriminate between two major types of lipids, sphingolipids and phospholipids, but were broadly cross-reactive towards diverse phospholipids including phosphatidylcholine, phosphatidylinositol and phosphatidylethanolamine. The crystal structure of a representative TCR bound to CD1b-phosphatidylcholine provides a molecular mechanism for this promiscuous recognition. We observe a lateral escape channel in the TCR, which shunted phospholipid head groups sideways along the CD1b-TCR interface, without contacting the TCR. Instead the TCR recognition site involved the neck region phosphate that is common to all major self-phospholipids but absent in sphingolipids. Whereas prior studies have focused on foreign lipids or rare self-lipids, we define a new molecular mechanism of promiscuous recognition of common self-phospholipids including those that are known targets in human autoimmune disease.
- Infection and Immunity Program and Department of Biochemistry and Molecular Biology, Biomedicine Discovery Institute, Monash University, Clayton, VIC, 3800, Australia.
Organizational Affiliation: