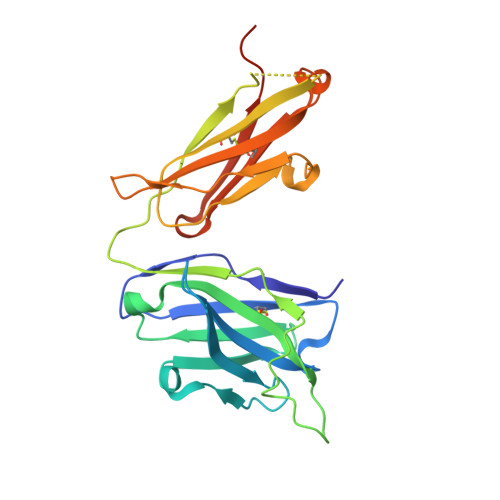
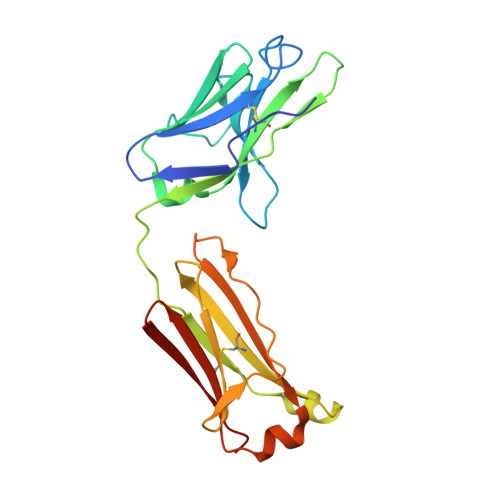
HIV envelope V3 region mimic embodies key features of a broadly neutralizing antibody lineage epitope.
Fera, D., Lee, M.S., Wiehe, K., Meyerhoff, R.R., Piai, A., Bonsignori, M., Aussedat, B., Walkowicz, W.E., Ton, T., Zhou, J.O., Danishefsky, S., Haynes, B.F., Harrison, S.C.(2018) Nat Commun 9: 1111-1111
- PubMed: 29549260 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-018-03565-6
- Primary Citation Related Structures:
6CBJ, 6CBP - PubMed Abstract:
HIV-1 envelope (Env) mimetics are candidate components of prophylactic vaccines and potential therapeutics. Here we use a synthetic V3-glycopeptide ("Man 9 -V3") for structural studies of an HIV Env third variable loop (V3)-glycan directed, broadly neutralizing antibody (bnAb) lineage ("DH270"), to visualize the epitope on Env and to study how affinity maturation of the lineage proceeded. Unlike many previous V3 mimetics, Man 9 -V3 encompasses two key features of the V3 region recognized by V3-glycan bnAbs-the conserved GDIR motif and the N332 glycan. In our structure of an antibody fragment of a lineage member, DH270.6, in complex with the V3 glycopeptide, the conformation of the antibody-bound glycopeptide conforms closely to that of the corresponding segment in an intact HIV-1 Env trimer. An additional structure identifies roles for two critical mutations in the development of breadth. The results suggest a strategy for use of a V3 glycopeptide as a vaccine immunogen.
- Laboratory of Molecular Medicine, Boston Children's Hospital, Harvard Medical School, Boston, MA, 02115, USA. dfera1@swarthmore.edu.
Organizational Affiliation: