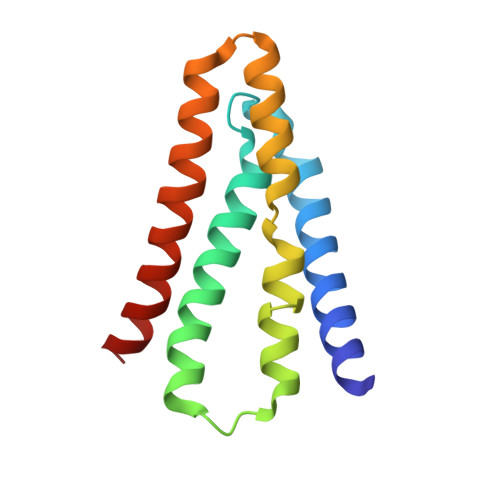
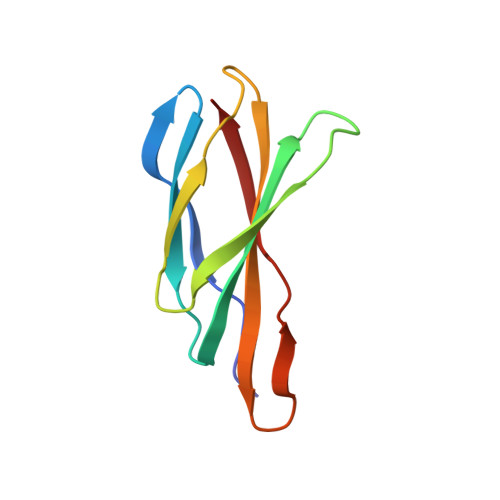
Molecular Interactions between a Fluoride Ion Channel and Synthetic Protein Blockers.
Turman, D.L., Cheloff, A.Z., Corrado, A.D., Nathanson, J.T., Miller, C.(2018) Biochemistry 57: 1212-1218
- PubMed: 29393634 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acs.biochem.7b01272
- Primary Citation Related Structures:
6BX4, 6BX5 - PubMed Abstract:
Fluoride ion channels of the Fluc family selectively export F - ions to rescue unicellular organisms from acute F - toxicity. Crystal structures of bacterial Fluc channels in complex with synthetic monobodies, fibronectin-derived soluble β-sandwich fold proteins, show 2-fold symmetric homodimers with an antiparallel transmembrane topology. Monobodies also block Fluc F - current via a pore blocking mechanism. However, little is known about the energetic contributions of individual monobody residues to the affinity of the monobody-channel complex or whether the structural paratope corresponds to functional reality. This study seeks to structurally identify and compare residues interacting with Fluc between two highly similar monobodies and subjects them to mutagenesis and functional measurements of equilibrium affinities via a fluorescence anisotropy binding assay to determine their energetic contributions. The results indicate that the functional and structural paratopes strongly agree and that many Tyr residues at the interface, while playing a key role in affinity, can be substituted with Phe and Trp without large disruptions.
- Department of Biochemistry, Howard Hughes Medical Institute, Brandeis University , 415 South Street, Waltham, Massachusetts 02453, United States.
Organizational Affiliation: