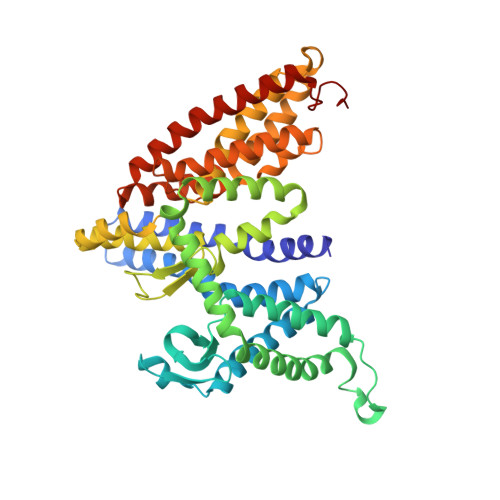
Structure of an EIIC sugar transporter trapped in an inward-facing conformation.
Ren, Z., Lee, J., Moosa, M.M., Nian, Y., Hu, L., Xu, Z., McCoy, J.G., Ferreon, A.C.M., Im, W., Zhou, M.(2018) Proc Natl Acad Sci U S A 115: 5962-5967
- PubMed: 29784777 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1800647115
- Primary Citation Related Structures:
6BVG - PubMed Abstract:
The phosphoenolpyruvate-dependent phosphotransferase system (PTS) transports sugar into bacteria and phosphorylates the sugar for metabolic consumption. The PTS is important for the survival of bacteria and thus a potential target for antibiotics, but its mechanism of sugar uptake and phosphorylation remains unclear. The PTS is composed of multiple proteins, and the membrane-embedded Enzyme IIC (EIIC) component transports sugars across the membrane. Crystal structures of two members of the glucose superfamily of EIICs, bcChbC and bcMalT, were solved in the inward-facing and outward-facing conformations, and the structures suggest that sugar translocation could be achieved by movement of a structured domain that contains the sugar-binding site. However, different conformations have not been captured on the same transporter to allow precise description of the conformational changes. Here we present a crystal structure of bcMalT trapped in an inward-facing conformation by a mercury ion that bridges two strategically placed cysteine residues. The structure allows direct comparison of the outward- and inward-facing conformations and reveals a large rigid-body motion of the sugar-binding domain and other conformational changes that accompany the rigid-body motion. All-atom molecular dynamics simulations show that the inward-facing structure is stable with or without the cross-linking. The conformational changes were further validated by single-molecule Föster resonance energy transfer (smFRET). Combined, these results establish the elevator-type mechanism of transport in the glucose superfamily of EIIC transporters.
- Department of Biochemistry and Molecular Biology, Baylor College of Medicine, Houston, TX 77030.
Organizational Affiliation: