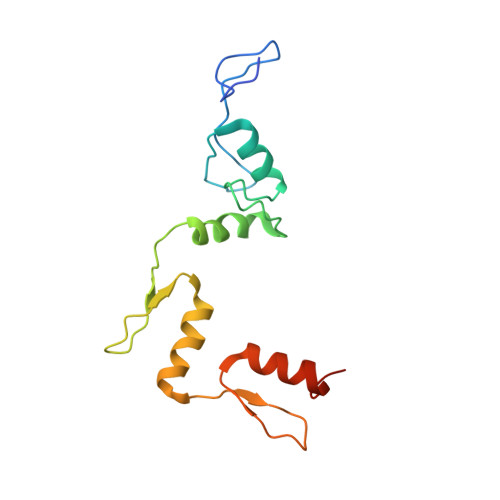
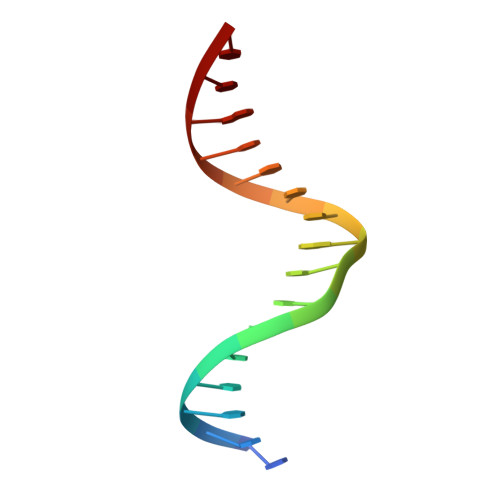
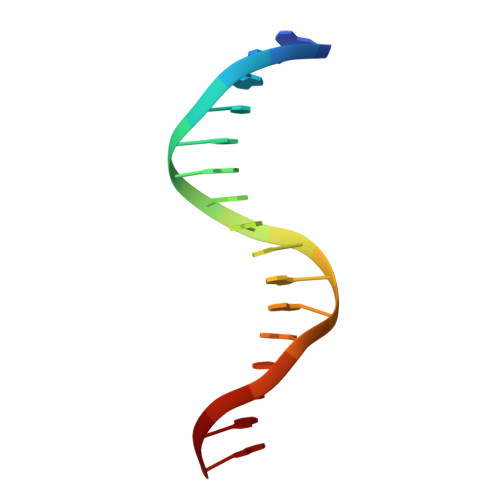
Crystal structures of REF6 and its complex with DNA reveal diverse recognition mechanisms.
Tian, Z., Li, X., Li, M., Wu, W., Zhang, M., Tang, C., Li, Z., Liu, Y., Chen, Z., Yang, M., Ma, L., Caba, C., Tong, Y., Lam, H.M., Dai, S., Chen, Z.(2020) Cell Discov 6: 17-17
- PubMed: 32257379 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41421-020-0150-6
- Primary Citation Related Structures:
6A57, 6A58, 6A59 - PubMed Abstract:
Relative of Early Flowing 6 (REF6) is a DNA-sequence-specific H3K27me3/2 demethylase that contains four zinc finger (ZnF) domains and targets several thousand genes in Arabidopsis thaliana . The ZnF domains are essential for binding target genes, but the structural basis remains unclear. Here, we determined crystal structures of the ZnF domains and REF6-DNA complex, revealing a unique REF6-family-specific half-cross-braced ZnF (RCZ) domain and two C2H2-type ZnFs. DNA-binding induces a profound conformational change in the hinge region of REF6. Each REF6 recognizes six bases and DNA methylation reduces the binding affinity. Both the acidic region and basic region are important for the self-association of REF6. The REF6 DNA-binding affinity is determined by the sequence-dependent conformations of DNA and also the cooperativity in different target motifs. The conformational plasticity enables REF6 to function as a global transcriptional regulator that directly binds to many diverse genes, revealing the structural basis for the epigenetic modification recognition.
- 1State Key Laboratory of Agrobiotechnology and Beijing Advanced Innovation Center for Food Nutrition and Human Health, College of Biological Sciences, China Agricultural University, 100193 Beijing, China.
Organizational Affiliation: