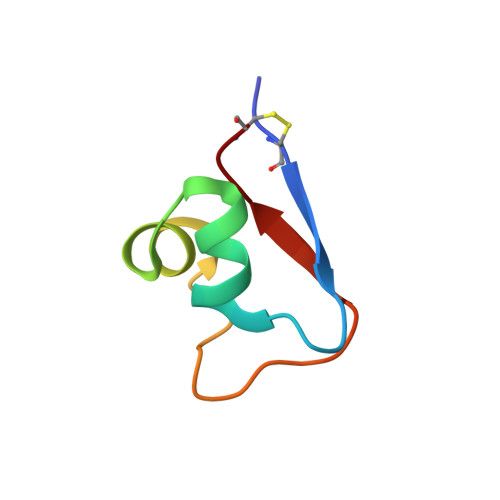
Structures and chitin-binding properties of two N-terminal lysin motifs (LysMs) found in a chitinase from Volvox carteri.
Kitaoku, Y., Nishimura, S., Hirono, T., Suginta, W., Ohnuma, T., Fukamizo, T.(2019) Glycobiology 29: 565-575
- PubMed: 30976779 Search on PubMed
- DOI: https://doi.org/10.1093/glycob/cwz024
- Primary Citation Related Structures:
5YZ6, 5YZK - PubMed Abstract:
Two N-terminal lysin motifs (LysMs) found in a chitinase from the green alga Volvox carteri (VcLysM1 and VcLysM2) were produced, and their structures and chitin-binding properties were characterized. The binding affinities of VcLysM1 toward chitin oligomers determined by isothermal titration calorimetry (ITC) were higher than those of VcLysM2 by 0.8-1.1 kcal/mol of ΔG°. Based on the NMR solution structures of the two LysMs, the differences in binding affinities were found to result from amino acid substitutions at the binding site. The NMR spectrum of a two-domain protein (VcLysM1+2), in which VcLysM1 and VcLysM2 are linked in tandem through a flexible linker, suggested that the individual domains of VcLysM1+2 independently fold and do not interact with each other. ITC analysis of chitin-oligomer binding revealed two different binding sites in VcLysM1+2, showing no cooperativity. The binding affinities of the VcLysM1 domain in VcLysM1+2 were lower than those of VcLysM1 alone, probably due to the flexible linker destabilizing the interaction between the chito-oligosaccahrides and VcLysM1 domain. Overall, two LysMs attached to the chitinase from the primitive plant species, V. carteri, were found to resemble bacterial LysMs reported thus far.
- Department of Advanced Bioscience, Kindai University, 3327-204 Nakamachi, Nara Japan.
Organizational Affiliation: