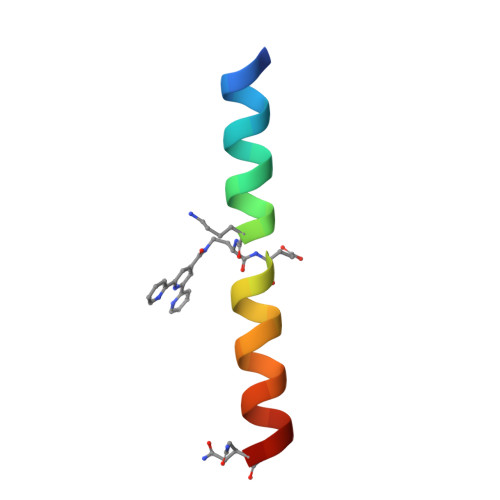
Supramolecular Metal-Coordination Polymers, Nets, and Frameworks from Synthetic Coiled-Coil Peptides.
Tavenor, N.A., Murnin, M.J., Horne, W.S.(2017) J Am Chem Soc 139: 2212-2215
- PubMed: 28161945 Search on PubMed
- DOI: https://doi.org/10.1021/jacs.7b00651
- Primary Citation Related Structures:
5U59, 5U5A, 5U5B, 5U5C - PubMed Abstract:
Metal coordination and peptide-directed self-assembly are two proven methods for creating defined supramolecular architectures. Here, we report a new class of crystalline materials based on coiled-coil peptides bearing unnatural metal-chelating terpyridine moieties. High-resolution structural characterization of lattices formed in the presence of Cu 2+ reveals a general assembly mechanism. Subtle sequence variation in the modular synthetic ligand dictates assembly morphology.
- Department of Chemistry, University of Pittsburgh , Pittsburgh, Pennsylvania 15260, United States.
Organizational Affiliation: