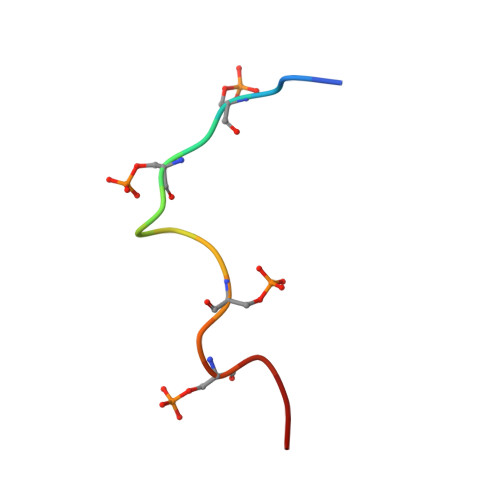
Structure and dynamics of the RNAPII CTDsome with Rtt103.
Jasnovidova, O., Klumpler, T., Kubicek, K., Kalynych, S., Plevka, P., Stefl, R.(2017) Proc Natl Acad Sci U S A 114: 11133-11138
- PubMed: 29073019 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1712450114
- Primary Citation Related Structures:
5M48, 5M9D - PubMed Abstract:
RNA polymerase II contains a long C-terminal domain (CTD) that regulates interactions at the site of transcription. The CTD architecture remains poorly understood due to its low sequence complexity, dynamic phosphorylation patterns, and structural variability. We used integrative structural biology to visualize the architecture of the CTD in complex with Rtt103, a 3'-end RNA-processing and transcription termination factor. Rtt103 forms homodimers via its long coiled-coil domain and associates densely on the repetitive sequence of the phosphorylated CTD via its N-terminal CTD-interacting domain. The CTD-Rtt103 association opens the compact random coil structure of the CTD, leading to a beads-on-a-string topology in which the long rod-shaped Rtt103 dimers define the topological and mobility restraints of the entire assembly. These findings underpin the importance of the structural plasticity of the CTD, which is templated by a particular set of CTD-binding proteins.
- CEITEC-Central European Institute of Technology, Masaryk University, CZ-62500 Brno, Czech Republic.
Organizational Affiliation: