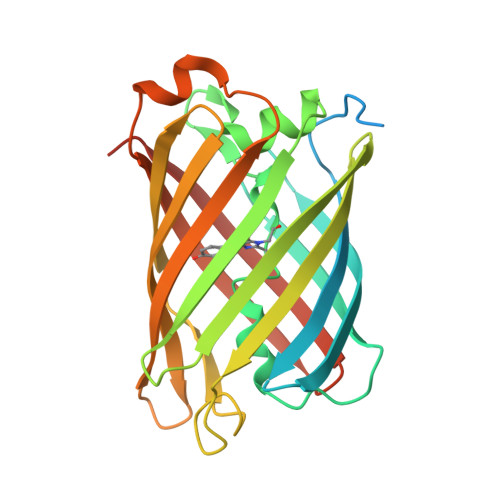
Structural analysis of the bright monomeric yellow-green fluorescent protein mNeonGreen obtained by directed evolution.
Clavel, D., Gotthard, G., von Stetten, D., De Sanctis, D., Pasquier, H., Lambert, G.G., Shaner, N.C., Royant, A.(2016) Acta Crystallogr D Struct Biol 72: 1298-1307
- PubMed: 27917830 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2059798316018623
- Primary Citation Related Structures:
5LTP, 5LTQ, 5LTR - PubMed Abstract:
Until recently, genes coding for homologues of the autofluorescent protein GFP had only been identified in marine organisms from the phyla Cnidaria and Arthropoda. New fluorescent-protein genes have now been found in the phylum Chordata, coding for particularly bright oligomeric fluorescent proteins such as the tetrameric yellow fluorescent protein lanYFP from Branchiostoma lanceolatum. A successful monomerization attempt led to the development of the bright yellow-green fluorescent protein mNeonGreen. The structures of lanYFP and mNeonGreen have been determined and compared in order to rationalize the directed evolution process leading from a bright, tetrameric to a still bright, monomeric fluorescent protein. An unusual discolouration of crystals of mNeonGreen was observed after X-ray data collection, which was investigated using a combination of X-ray crystallography and UV-visible absorption and Raman spectroscopies, revealing the effects of specific radiation damage in the chromophore cavity. It is shown that X-rays rapidly lead to the protonation of the phenolate O atom of the chromophore and to the loss of its planarity at the methylene bridge.
- Université Grenoble Alpes, Institut de Biologie Structurale (IBS), F-38044 Grenoble, France.
Organizational Affiliation: