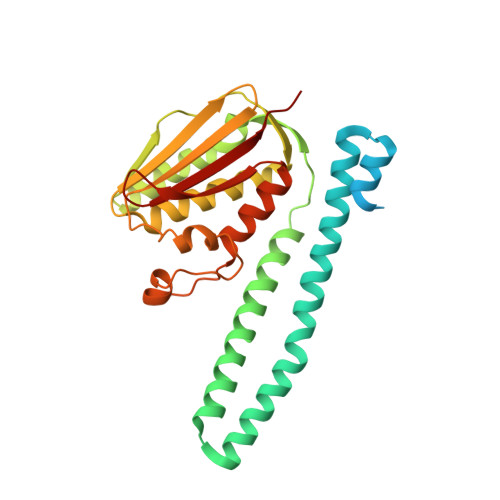
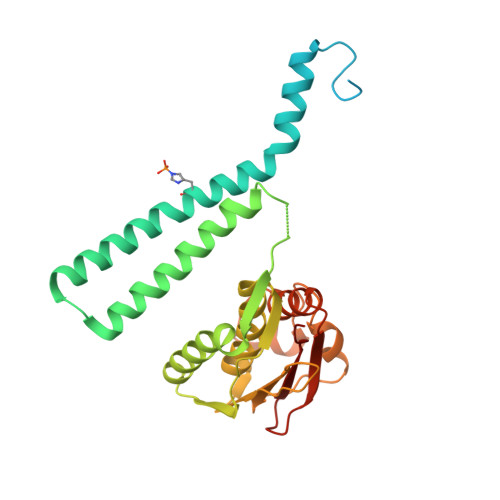
Structural Coupling between Autokinase and Phosphotransferase Reactions in a Bacterial Histidine Kinase.
Mechaly, A.E., Soto Diaz, S., Sassoon, N., Buschiazzo, A., Betton, J.M., Alzari, P.M.(2017) Structure 25: 939-944.e3
- PubMed: 28552574 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2017.04.011
- Primary Citation Related Structures:
4UHJ, 4UHK, 5LFK - PubMed Abstract:
Bacterial two-component systems consist of a sensor histidine kinase (HK) and a response regulator (RR). HKs are homodimers that catalyze the autophosphorylation of a histidine residue and the subsequent phosphoryl transfer to its RR partner, triggering an adaptive response. How the HK autokinase and phosphotransferase activities are coordinated remains unclear. Here, we report X-ray structures of the prototypical HK CpxA trapped as a hemi-phosphorylated dimer, and of the receiver domain from the RR partner, CpxR. Our results reveal that the two catalytic reactions can occur simultaneously, one in each protomer of the asymmetric CpxA dimer. Furthermore, the increase of autokinase activity in the presence of phosphotransfer-impaired CpxR put forward the idea of an allosteric switching mechanism, according to which CpxR binding to one CpxA protomer triggers autophosphorylation in the second protomer. The ensuing dynamical model provides a mechanistic explanation of how HKs can efficiently orchestrate two catalytic reactions involving large-scale protein motions.
- Institut Pasteur, Unité de Microbiologie Structurale, CNRS UMR 3528 & Université Paris Diderot, Sorbonne Paris Cité, 25 rue du Dr. Roux, 75724, Paris Cedex 15, France. Electronic address: aemechaly@pasteur.edu.uy.
Organizational Affiliation: