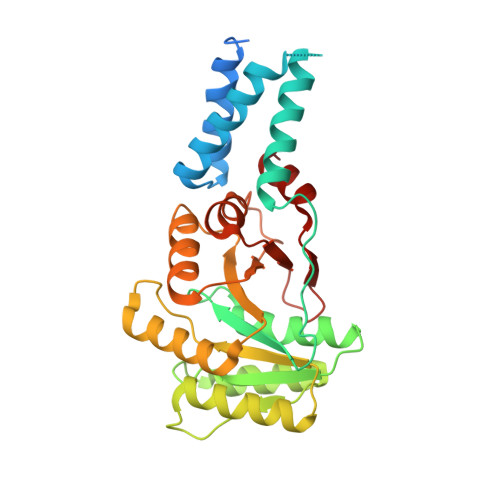
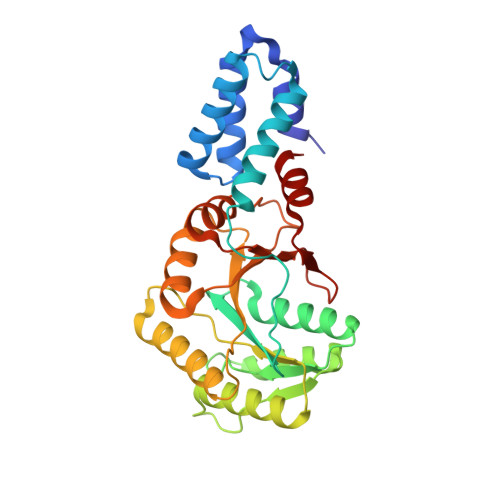
Structural Basis for Conserved Regulation and Adaptation of the Signal Recognition Particle Targeting Complex.
Wild, K., Bange, G., Motiejunas, D., Kribelbauer, J., Hendricks, A., Segnitz, B., Wade, R.C., Sinning, I.(2016) J Mol Biology 428: 2880-2897
- PubMed: 27241309 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2016.05.015
- Primary Citation Related Structures:
5L3Q, 5L3R, 5L3S, 5L3V, 5L3W - PubMed Abstract:
The signal recognition particle (SRP) is a ribonucleoprotein complex with a key role in targeting and insertion of membrane proteins. The two SRP GTPases, SRP54 (Ffh in bacteria) and FtsY (SRα in eukaryotes), form the core of the targeting complex (TC) regulating the SRP cycle. The architecture of the TC and its stimulation by RNA has been described for the bacterial SRP system while this information is lacking for other domains of life. Here, we present the crystal structures of the GTPase heterodimers of archaeal (Sulfolobus solfataricus), eukaryotic (Homo sapiens), and chloroplast (Arabidopsis thaliana) SRP systems. The comprehensive structural comparison combined with Brownian dynamics simulations of TC formation allows for the description of the general blueprint and of specific adaptations of the quasi-symmetric heterodimer. Our work defines conserved external nucleotide-binding sites for SRP GTPase activation by RNA. Structural analyses of the GDP-bound, post-hydrolysis states reveal a conserved, magnesium-sensitive switch within the I-box. Overall, we provide a general model for SRP cycle regulation by RNA.
- Heidelberg University Biochemistry Center (BZH), INF 328, D-69120 Heidelberg, Germany.
Organizational Affiliation: