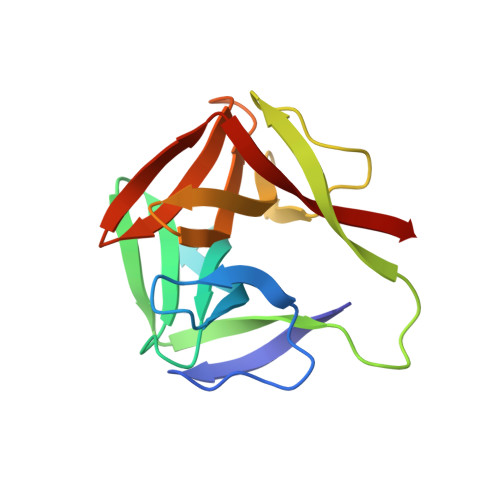
Identification, Characterization, and X-ray Crystallographic Analysis of a Novel Type of Mannose-Specific Lectin CGL1 from the Pacific Oyster Crassostrea gigas.
Unno, H., Matsuyama, K., Tsuji, Y., Goda, S., Hiemori, K., Tateno, H., Hirabayashi, J., Hatakeyama, T.(2016) Sci Rep 6: 29135-29135
- PubMed: 27377186 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/srep29135
- Primary Citation Related Structures:
5ID8, 5IDA, 5IDB - PubMed Abstract:
A novel mannose-specific lectin, named CGL1 (15.5 kDa), was isolated from the oyster Crassostrea gigas. Characterization of CGL1 involved isothermal titration calorimetry (ITC), glycoconjugate microarray, and frontal affinity chromatography (FAC). This analysis revealed that CGL1 has strict specificity for the mannose monomer and for high mannose-type N-glycans (HMTGs). Primary structure of CGL1 did not show any homology with known lectins but did show homology with proteins of the natterin family. Crystal structure of the CGL1 revealed a unique homodimer in which each protomer was composed of 2 domains related by a pseudo two-fold axis. Complex structures of CGL1 with mannose molecules showed that residues have 8 hydrogen bond interactions with O1, O2, O3, O4, and O5 hydroxyl groups of mannose. The complex interactions that are not observed with other mannose-binding lectins revealed the structural basis for the strict specificity for mannose. These characteristics of CGL1 may be helpful as a research tool and for clinical applications.
- Graduate School of Engineering, Nagasaki University, 1-14 Bunkyo-machi, Nagasaki 852-8521, Japan.
Organizational Affiliation: