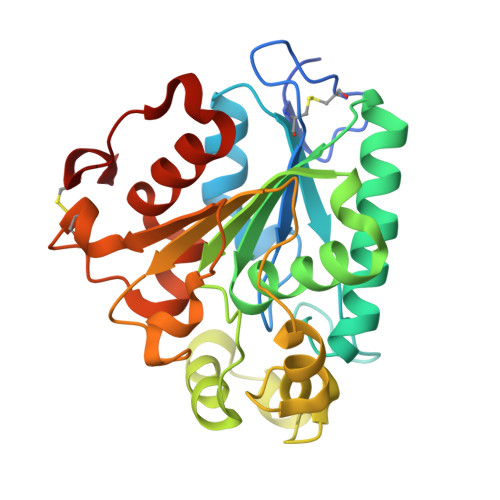
Crystal structure of a lipase from Streptomyces sp. strain W007 - implications for thermostability and regiospecificity
Zhao, Z., Hou, S., Lan, D., Wang, X., Liu, J., Khan, F.I., Wang, Y.(2017) FEBS J 284: 3506-3519
- PubMed: 28857479 Search on PubMed
- DOI: https://doi.org/10.1111/febs.14211
- Primary Citation Related Structures:
5H6B, 5H6G - PubMed Abstract:
MAS1 from marine Streptomyces sp. strain W007 belongs to the bacterial lipase I.7 subfamily and is characterized as a thermostable and nonregiospecific lipase. To shed light on the catalytic mechanism of MAS1, we determined its crystal structure with closed conformation in two crystal forms at 2.3 Å resolution. MAS1 adopts the canonical α/β hydrolase core fold with its catalytic triad being formed by S109, D200 and H232. Structural analysis and biochemical assays revealed that disulfide bonds and salt bridges play a vital role in the thermostability of MAS1. In addition, we discovered that the replacement of H108 with a tryptophan converts MAS1 from a nonregiospecific to an sn-1,3-specific lipase, suggesting the functional importance of the second position from the conserved pentapeptide motif in defining the regiospecificity of MAS1. Our present study provides insights into the molecular basis for the thermostability and regiospecificity of MAS1, and it may aid in the rational design of thermostable or regiospecific lipases for potential industrial applications. Structural data are available in the Protein Data Bank database under the accession numbers 5H6B and 5H6G.
- School of Bioscience and Bioengineering, South China University of Technology, Guangzhou, China.
Organizational Affiliation: