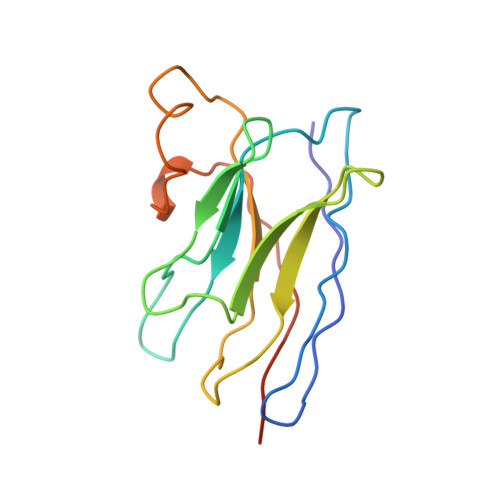
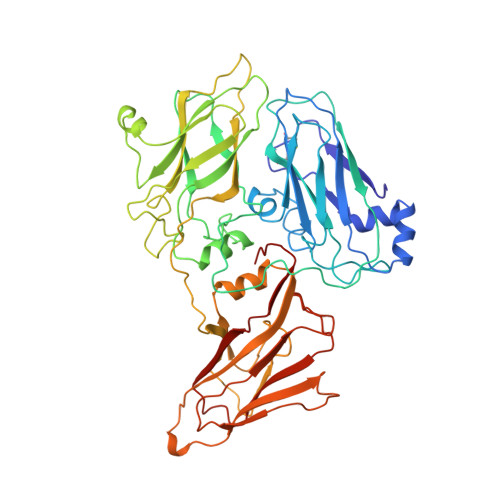
Structural basis of nanobody recognition of grapevine fanleaf virus and of virus resistance loss.
Orlov, I., Hemmer, C., Ackerer, L., Lorber, B., Ghannam, A., Poignavent, V., Hleibieh, K., Sauter, C., Schmitt-Keichinger, C., Belval, L., Hily, J.M., Marmonier, A., Komar, V., Gersch, S., Schellenberger, P., Bron, P., Vigne, E., Muyldermans, S., Lemaire, O., Demangeat, G., Ritzenthaler, C., Klaholz, B.P.(2020) Proc Natl Acad Sci U S A
- PubMed: 32371486 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1913681117
- Primary Citation Related Structures:
5FOJ - PubMed Abstract:
Grapevine fanleaf virus (GFLV) is a picorna-like plant virus transmitted by nematodes that affects vineyards worldwide. Nanobody (Nb)-mediated resistance against GFLV has been created recently, and shown to be highly effective in plants, including grapevine, but the underlying mechanism is unknown. Here we present the high-resolution cryo electron microscopy structure of the GFLV-Nb23 complex, which provides the basis for molecular recognition by the Nb. The structure reveals a composite binding site bridging over three domains of one capsid protein (CP) monomer. The structure provides a precise mapping of the Nb23 epitope on the GFLV capsid in which the antigen loop is accommodated through an induced-fit mechanism. Moreover, we uncover and characterize several resistance-breaking GFLV isolates with amino acids mapping within this epitope, including C-terminal extensions of the CP, which would sterically interfere with Nb binding. Escape variants with such extended CP fail to be transmitted by nematodes linking Nb-mediated resistance to vector transmission. Together, these data provide insights into the molecular mechanism of Nb23-mediated recognition of GFLV and of virus resistance loss.
- Centre for Integrative Biology (CBI), Department of Integrated Structural Biology, IGBMC, Université de Strasbourg, 67404 Illkirch, France.
Organizational Affiliation: