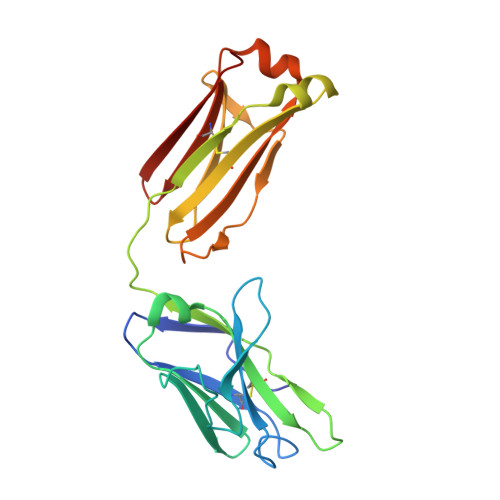
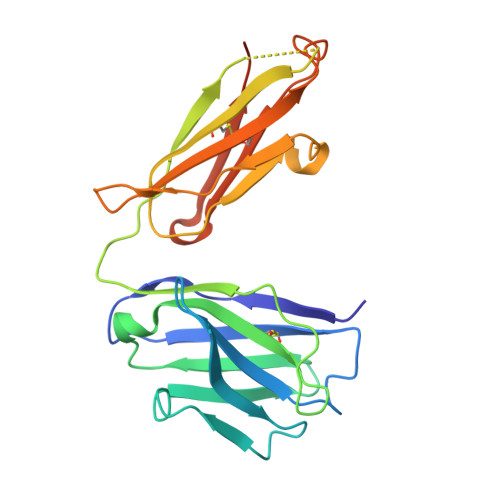
Structural Constraints of Vaccine-Induced Tier-2 Autologous HIV Neutralizing Antibodies Targeting the Receptor-Binding Site.
Bradley, T., Fera, D., Bhiman, J., Eslamizar, L., Lu, X., Anasti, K., Zhang, R., Sutherland, L.L., Scearce, R.M., Bowman, C.M., Stolarchuk, C., Lloyd, K.E., Parks, R., Eaton, A., Foulger, A., Nie, X., Karim, S.S., Barnett, S., Kelsoe, G., Kepler, T.B., Alam, S.M., Montefiori, D.C., Moody, M.A., Liao, H.X., Morris, L., Santra, S., Harrison, S.C., Haynes, B.F.(2016) Cell Rep 14: 43-54
- PubMed: 26725118 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.celrep.2015.12.017
- Primary Citation Related Structures:
5F6H, 5F6I, 5F6J - PubMed Abstract:
Antibodies that neutralize autologous transmitted/founder (TF) HIV occur in most HIV-infected individuals and can evolve to neutralization breadth. Autologous neutralizing antibodies (nAbs) against neutralization-resistant (Tier-2) viruses are rarely induced by vaccination. Whereas broadly neutralizing antibody (bnAb)-HIV-Envelope structures have been defined, the structures of autologous nAbs have not. Here, we show that immunization with TF mutant Envs gp140 oligomers induced high-titer, V5-dependent plasma neutralization for a Tier-2 autologous TF evolved mutant virus. Structural analysis of autologous nAb DH427 revealed binding to V5, demonstrating the source of narrow nAb specificity and explaining the failure to acquire breadth. Thus, oligomeric TF Envs can elicit autologous nAbs to Tier-2 HIVs, but induction of bnAbs will require targeting of precursors of B cell lineages that can mature to heterologous neutralization.
- Duke Human Vaccine Institute, Departments of Medicine, Surgery and Immunology, Duke University School of Medicine, Durham, NC 27710, USA. Electronic address: todd.bradley@duke.edu.
Organizational Affiliation: