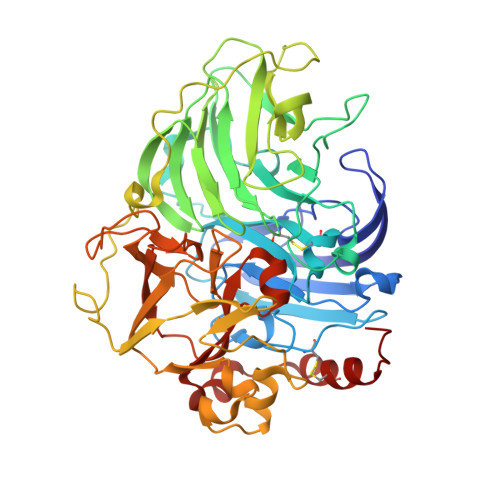
Structure-function study of two new middle-redox potential laccases from basidiomycetes Antrodiella faginea and Steccherinum murashkinskyi.
Glazunova, O.A., Polyakov, K.M., Moiseenko, K.V., Kurzeev, S.A., Fedorova, T.V.(2018) Int J Biol Macromol 118: 406-418
- PubMed: 29890251 Search on PubMed
- DOI: https://doi.org/10.1016/j.ijbiomac.2018.06.038
- Primary Citation Related Structures:
5E9N, 5EHF - PubMed Abstract:
Laccases are multicopper oxidases that catalyze oxidation of a wide range of organic and inorganic substrates accompanied by the reduction of dioxygen to water. The physicochemical and catalytic properties of two new fungal laccases from basidiomycetes Antrodiella faginea (AfL) and Steccherinum murashkinskyi (SmL) with middle redox potential of the T1 copper site were studied. The X-ray structures of AfL and SmL were solved at 1.75 Å and 0.95 Å, respectively. The oxidized state of copper ions in the active site was observed in AfL structure, while the mixture of oxidized and reduced states was observed in SmL structure. These oxidized and reduced states relate to the position of copper ions, their coordination, and nature and position of oxygen ligands. Comparative analysis of the T1 site environment of laccases with known structure allowed us to highlight the six types of the secondary coordination sphere of the T1 copper. The solvent accessible surface area of the conservative region of the secondary coordination sphere of the T1 copper correlates with its the redox potential. It was shown that the laccase classification by the structure of the T1 copper secondary coordination sphere is in agreement to ecophysiological behavior of laccase producing fungi.
- A. N. Bach Institute of Biochemistry, Research Center of Biotechnology, Russian Academy of Sciences, Leninsky Ave. 33/2, Moscow 119071, Russian Federation. Electronic address: olga.a.glas@gmail.com.
Organizational Affiliation: