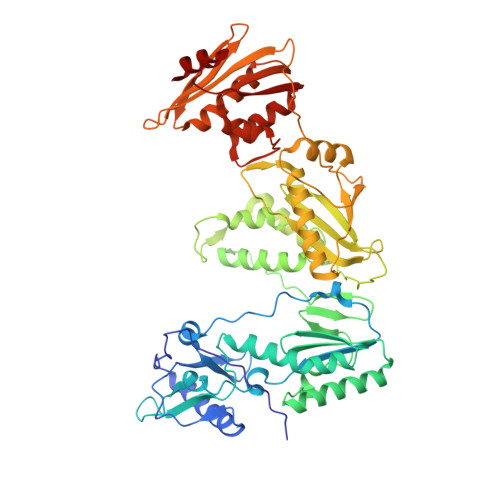
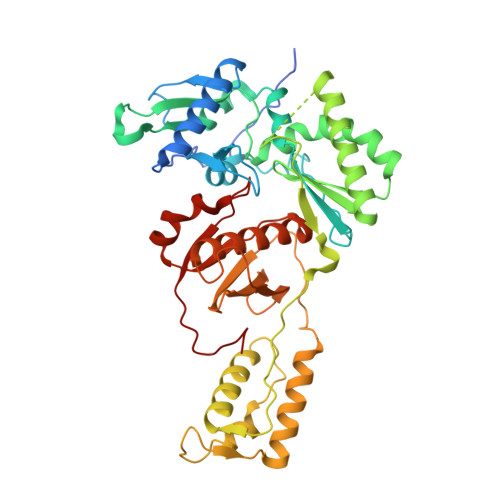
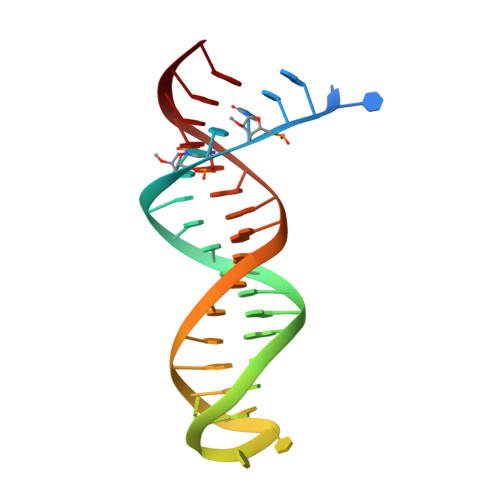
Structure of HIV-1 reverse transcriptase bound to a novel 38-mer hairpin template-primer DNA aptamer.
Miller, M.T., Tuske, S., Das, K., DeStefano, J.J., Arnold, E.(2016) Protein Sci 25: 46-55
- PubMed: 26296781 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.2776
- Primary Citation Related Structures:
5D3G - PubMed Abstract:
The development of a modified DNA aptamer that binds HIV-1 reverse transcriptase (RT) with ultra-high affinity has enabled the X-ray structure determination of an HIV-1 RT-DNA complex to 2.3 Å resolution without the need for an antibody Fab fragment or RT-DNA cross-linking. The 38-mer hairpin-DNA aptamer has a 15 base-pair duplex, a three-deoxythymidine hairpin loop, and a five-nucleotide 5'-overhang. The aptamer binds RT in a template-primer configuration with the 3'-end positioned at the polymerase active site and has 2'-O-methyl modifications at the second and fourth duplex template nucleotides that interact with the p66 fingers and palm subdomains. This structure represents the highest resolution RT-nucleic acid structure to date. The RT-aptamer complex is catalytically active and can serve as a platform for studying fundamental RT mechanisms and for development of anti-HIV inhibitors through fragment screening and other approaches. Additionally, the structure allows for a detailed look at a unique aptamer design and provides the molecular basis for its remarkably high affinity for RT.
- Center for Advanced Biotechnology and Medicine, Department of Chemistry and Chemical Biology, Rutgers University, Piscataway, New Jersey, 08854.
Organizational Affiliation: