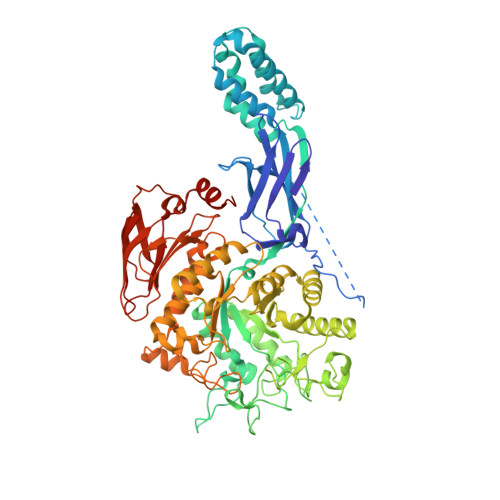
Structure of Mycobacterium thermoresistibile GlgE defines novel conformational states that contribute to the catalytic mechanism.
Mendes, V., Blaszczyk, M., Maranha, A., Empadinhas, N., Blundell, T.L.(2015) Sci Rep 5: 17144-17144
- PubMed: 26616850 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/srep17144
- Primary Citation Related Structures:
5CGM, 5CIM, 5CJ5 - PubMed Abstract:
GlgE, an enzyme of the pathway that converts trehalose to α-glucans, is essential for Mycobacterium tuberculosis. Inhibition of GlgE, which transfers maltose from a maltose-1-phosphate donor to α-glucan/maltooligosaccharide chain acceptor, leads to a toxic accumulation of maltose-1-phosphate that culminates in cellular death. Here we describe the first high-resolution mycobacterial GlgE structure from Mycobacterium thermoresistibile at 1.96 Å. We show that the structure resembles that of M. tuberculosis and Streptomyces coelicolor GlgEs, reported before, with each protomer in the homodimer comprising five domains. However, in M. thermoresistibile GlgE we observe several conformational states of the S domain and provide evidence that its high flexibility is important for enzyme activity. The structures here reported shed further light on the interactions between the N-terminal domains and the catalytic domains of opposing chains and how they contribute to the catalytic reaction. Importantly this work identifies a useful surrogate system to aid the development of GlgE inhibitors against opportunistic and pathogenic mycobacteria.
- Department of Biochemistry, University of Cambridge, Cambridge CB2 1GA, UK.
Organizational Affiliation: