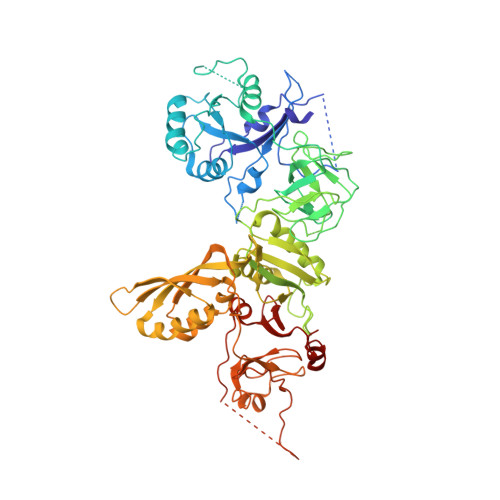
Structure of Bipa in GTP Form Bound to the Ratcheted Ribosome.
Kumar, V., Chen, Y., Ero, R., Ahmed, T., Tan, J., Li, Z., Wong, A.S.W., Bhushan, S., Gao, Y.(2015) Proc Natl Acad Sci U S A 112: 10944
- PubMed: 26283392 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1513216112
- Primary Citation Related Structures:
5A9V, 5A9W, 5A9X, 5A9Y, 5A9Z, 5AA0 - PubMed Abstract:
BPI-inducible protein A (BipA) is a member of the family of ribosome-dependent translational GTPase (trGTPase) factors along with elongation factors G and 4 (EF-G and EF4). Despite being highly conserved in bacteria and playing a critical role in coordinating cellular responses to environmental changes, its structures (isolated and ribosome bound) remain elusive. Here, we present the crystal structures of apo form and GTP analog, GDP, and guanosine-3',5'-bisdiphosphate (ppGpp)-bound BipA. In addition to having a distinctive domain arrangement, the C-terminal domain of BipA has a unique fold. Furthermore, we report the cryo-electron microscopy structure of BipA bound to the ribosome in its active GTP form and elucidate the unique structural attributes of BipA interactions with the ribosome and A-site tRNA in the light of its possible function in regulating translation.
- Institute of Molecular and Cell Biology, A*STAR, 138673, Singapore; School of Biological Sciences, Nanyang Technological University, 637551, Singapore.
Organizational Affiliation: