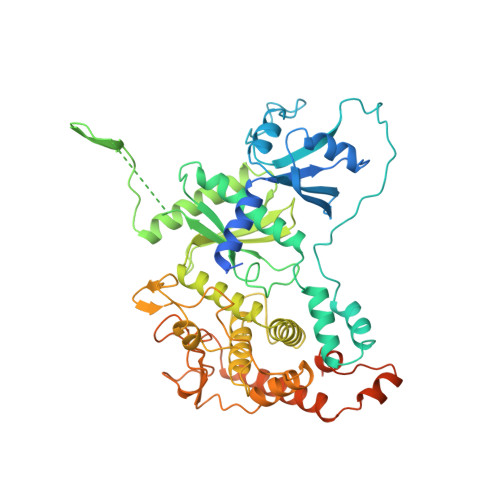
Structure of Mitochondrial Poly(A) RNA Polymerase Reveals the Structural Basis for Dimerization, ATP Selectivity and the Spax4 Disease Phenotype.
Lapkouski, M., Hallberg, B.M.(2015) Nucleic Acids Res 43: 9065
- PubMed: 26319014 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkv861
- Primary Citation Related Structures:
5A2V, 5A2W, 5A2X, 5A2Y, 5A2Z, 5A30 - PubMed Abstract:
Polyadenylation, performed by poly(A) polymerases (PAPs), is a ubiquitous post-transcriptional modification that plays key roles in multiple aspects of RNA metabolism. Although cytoplasmic and nuclear PAPs have been studied extensively, the mechanism by which mitochondrial PAP (mtPAP) selects adenosine triphosphate over other nucleotides is unknown. Furthermore, mtPAP is unique because it acts as a dimer. However, mtPAP's dimerization requirement remains enigmatic. Here, we show the structural basis for mtPAP's nucleotide selectivity, dimerization and catalysis. Our structures reveal an intricate dimerization interface that features an RNA-recognition module formed through strand complementation. Further, we propose the structural basis for the N478D mutation that drastically reduces the length of poly(A) tails on mitochondrial mRNAs in patients with spastic ataxia 4 (SPAX4), a severe and progressive neurodegenerative disease.
- Department of Cell and Molecular Biology, Karolinska Institutet, 171 77 Stockholm, Sweden Röntgen-Ångström-Cluster, Karolinska Institutet Outstation, Centre for Structural Systems Biology, DESY-Campus, 22607 Hamburg, Germany.
Organizational Affiliation: