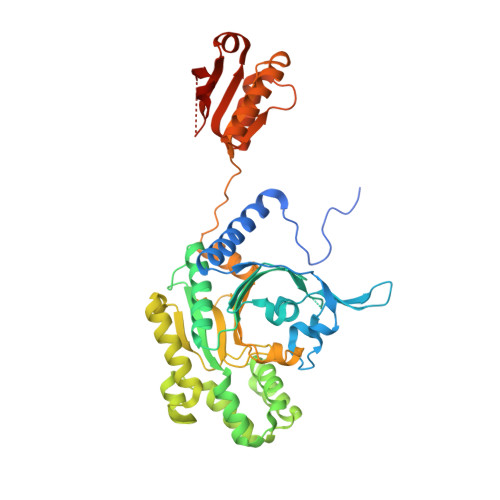
A binding hotspot in Trypanosoma cruzi histidyl-tRNA synthetase revealed by fragment-based crystallographic cocktail screens.
Koh, C.Y., Siddaramaiah, L.K., Ranade, R.M., Nguyen, J., Jian, T., Zhang, Z., Gillespie, J.R., Buckner, F.S., Verlinde, C.L., Fan, E., Hol, W.G.(2015) Acta Crystallogr D Biol Crystallogr 71: 1684-1698
- PubMed: 26249349 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S1399004715007683
- Primary Citation Related Structures:
4YP0, 4YPF, 4YRC, 4YRE, 4YRF, 4YRG, 4YRI, 4YRJ, 4YRK, 4YRL, 4YRM, 4YRN, 4YRO, 4YRP, 4YRQ, 4YRR, 4YRS, 4YRT - PubMed Abstract:
American trypanosomiasis, commonly known as Chagas disease, is a neglected tropical disease caused by the protozoan parasite Trypanosoma cruzi. The chronic form of the infection often causes debilitating morbidity and mortality. However, the current treatment for the disease is typically inadequate owing to drug toxicity and poor efficacy, necessitating a continual effort to discover and develop new antiparasitic therapeutic agents. The structure of T. cruzi histidyl-tRNA synthetase (HisRS), a validated drug target, has previously been reported. Based on this structure and those of human cytosolic HisRS, opportunities for the development of specific inhibitors were identified. Here, efforts are reported to identify small molecules that bind to T. cruzi HisRS through fragment-based crystallographic screening in order to arrive at chemical starting points for the development of specific inhibitors. T. cruzi HisRS was soaked into 68 different cocktails from the Medical Structural Genomics of Pathogenic Protozoa (MSGPP) fragment library and diffraction data were collected to identify bound fragments after soaking. A total of 15 fragments were identified, all bound to the same site on the protein, revealing a fragment-binding hotspot adjacent to the ATP-binding pocket. On the basis of the initial hits, the design of reactive fragments targeting the hotspot which would be simultaneously covalently linked to a cysteine residue present only in trypanosomatid HisRS was initiated. Inhibition of T. cruzi HisRS was observed with the resultant reactive fragments and the anticipated binding mode was confirmed crystallographically. These results form a platform for the development of future generations of selective inhibitors for trypanosomatid HisRS.
- Department of Biochemistry, University of Washington, Seattle, WA 98195, USA.
Organizational Affiliation: