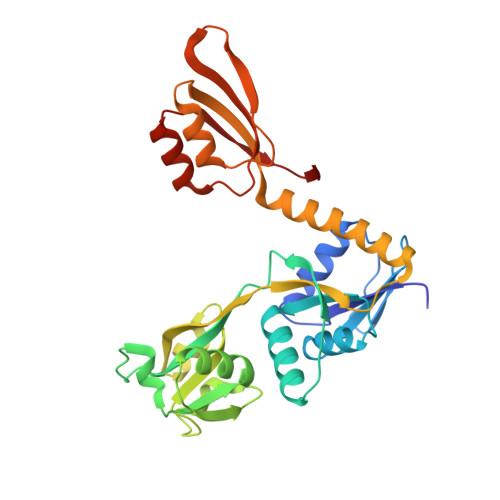
Campylobacter jejuni adenosine triphosphate phosphoribosyltransferase is an active hexamer that is allosterically controlled by the twisting of a regulatory tail.
Mittelstadt, G., Moggre, G.J., Panjikar, S., Nazmi, A.R., Parker, E.J.(2016) Protein Sci 25: 1492-1506
- PubMed: 27191057 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.2948
- Primary Citation Related Structures:
4YB5, 4YB6, 4YB7 - PubMed Abstract:
Adenosine triphosphate phosphoribosyltransferase (ATP-PRT) catalyzes the first committed step of the histidine biosynthesis in plants and microorganisms. Here, we present the functional and structural characterization of the ATP-PRT from the pathogenic ε-proteobacteria Campylobacter jejuni (CjeATP-PRT). This enzyme is a member of the long form (HisGL ) ATP-PRT and is allosterically inhibited by histidine, which binds to a remote regulatory domain, and competitively inhibited by AMP. In the crystalline form, CjeATP-PRT was found to adopt two distinctly different hexameric conformations, with an open homohexameric structure observed in the presence of substrate ATP, and a more compact closed form present when inhibitor histidine is bound. CjeATP-PRT was observed to adopt only a hexameric quaternary structure in solution, contradicting previous hypotheses favoring an allosteric mechanism driven by an oligomer equilibrium. Instead, this study supports the conclusion that the ATP-PRT long form hexamer is the active species; the tightening of this structure in response to remote histidine binding results in an inhibited enzyme.
- Maurice Wilkins Centre, Biomolecular Interaction Centre, Christchurch, 8140, New Zealand.
Organizational Affiliation: