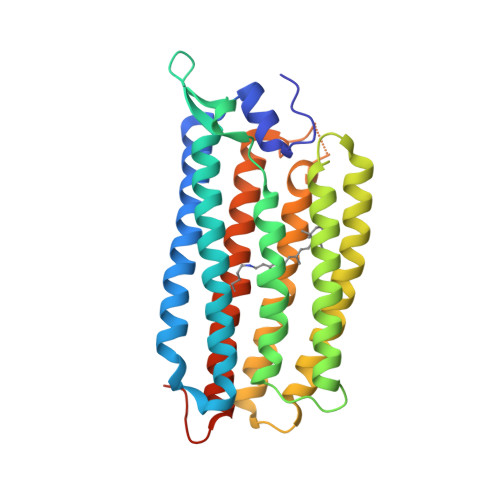
Crystal structure of a light-driven sodium pump.
Gushchin, I., Shevchenko, V., Polovinkin, V., Kovalev, K., Alekseev, A., Round, E., Borshchevskiy, V., Balandin, T., Popov, A., Gensch, T., Fahlke, C., Bamann, C., Willbold, D., Buldt, G., Bamberg, E., Gordeliy, V.(2015) Nat Struct Mol Biol 22: 390-395
- PubMed: 25849142 Search on PubMed
- DOI: https://doi.org/10.1038/nsmb.3002
- Primary Citation Related Structures:
4XTL, 4XTN, 4XTO - PubMed Abstract:
Recently, the first known light-driven sodium pumps, from the microbial rhodopsin family, were discovered. We have solved the structure of one of them, Krokinobacter eikastus rhodopsin 2 (KR2), in the monomeric blue state and in two pentameric red states, at resolutions of 1.45 Å and 2.2 and 2.8 Å, respectively. The structures reveal the ion-translocation pathway and show that the sodium ion is bound outside the protein at the oligomerization interface, that the ion-release cavity is capped by a unique N-terminal α-helix and that the ion-uptake cavity is unexpectedly large and open to the surface. Obstruction of the cavity with the mutation G263F imparts KR2 with the ability to pump potassium. These results pave the way for the understanding and rational design of cation pumps with new specific properties valuable for optogenetics.
- 1] Institut de Biologie Structurale, Université Grenoble Alpes, Grenoble, France. [2] Institut de Biologie Structurale, Centre National de la Recherche Scientifique, Grenoble, France. [3] Institut de Biologie Structurale, Commissariat à l'Énergie Atomique (CEA), Grenoble, France. [4] Laboratory for Advanced Studies of Membrane Proteins, Moscow Institute of Physics and Technology, Dolgoprudniy, Russia. [5] Institute of Complex Systems (ICS), ICS-6, Structural Biochemistry, Research Center Jülich, Jülich, Germany.
Organizational Affiliation: