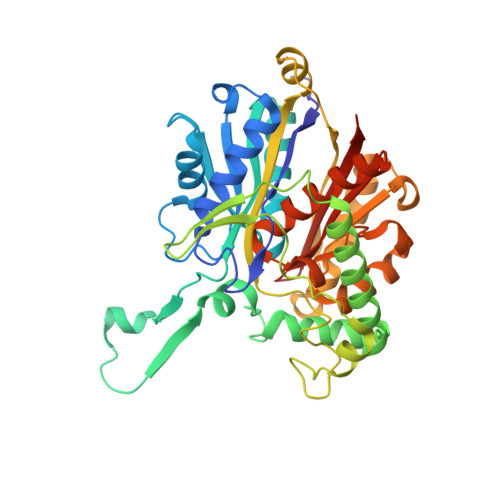
Redox-switch regulatory mechanism of thiolase from Clostridium acetobutylicum
Kim, S., Jang, Y.S., Ha, S.C., Ahn, J.W., Kim, E.J., Hong Lim, J., Cho, C., Shin Ryu, Y., Kuk Lee, S., Lee, S.Y., Kim, K.J.(2015) Nat Commun 6: 8410-8410
- PubMed: 26391388 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/ncomms9410
- Primary Citation Related Structures:
4WYR, 4WYS, 4XL2, 4XL3, 4XL4 - PubMed Abstract:
Thiolase is the first enzyme catalysing the condensation of two acetyl-coenzyme A (CoA) molecules to form acetoacetyl-CoA in a dedicated pathway towards the biosynthesis of n-butanol, an important solvent and biofuel. Here we elucidate the crystal structure of Clostridium acetobutylicum thiolase (CaTHL) in its reduced/oxidized states. CaTHL, unlike those from other aerobic bacteria such as Escherichia coli and Zoogloea ramegera, is regulated by the redox-switch modulation through reversible disulfide bond formation between two catalytic cysteine residues, Cys88 and Cys378. When CaTHL is overexpressed in wild-type C. acetobutylicum, butanol production is reduced due to the disturbance of acidogenic to solventogenic shift. The CaTHL(V77Q/N153Y/A286K) mutant, which is not able to form disulfide bonds, exhibits higher activity than wild-type CaTHL, and enhances butanol production upon overexpression. On the basis of these results, we suggest that CaTHL functions as a key enzyme in the regulation of the main metabolism of C. acetobutylicum through a redox-switch regulatory mechanism.
- School of Life Sciences, KNU Creative BioResearch Group, Kyungpook National University, Daegu 702-701, Korea.
Organizational Affiliation: