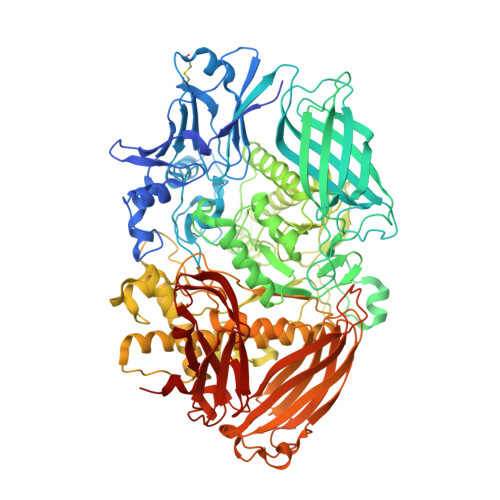
Insights Into the Structure and Function of Fungal Beta-Mannosidases from Glycoside Hydrolase Family 2 Based on Multiple Crystal Structures of the Trichoderma Harzianum Enzyme.
Nascimento, A.S., Muniz, J.R.C., Aparicio, R., Golubev, A.M., Polikarpov, I.(2014) FEBS J 281: 4165
- PubMed: 24975648 Search on PubMed
- DOI: https://doi.org/10.1111/febs.12894
- Primary Citation Related Structures:
4CVU, 4UOJ - PubMed Abstract:
Hemicellulose is an important part of the plant cell wall biomass, and is relevant to cellulosic ethanol technologies. β-Mannosidases are enzymes capable of cleaving nonreducing residues of β-d-mannose from β-d-mannosides and hemicellulose mannose-containing polysaccharides, such as mannans and galactomannans. β-Mannosidases are distributed between glycoside hydrolase (GH) families 1, 2, and 5, and only a handful of the enzymes have been structurally characterized to date. The only published X-ray structure of a GH family 2 mannosidase is that of the bacterial Bacteroides thetaiotaomicron enzyme. No structures of eukaryotic mannosidases of this family are currently available. To fill this gap, we set out to solve the structure of Trichoderma harzianum GH family 2 β-mannosidase and to refine it to 1.9-Å resolution. Structural comparisons of the T. harzianum GH2 β-mannosidase highlight similarities in its structural architecture with other members of GH family 2, reveal the molecular mechanism of β-mannoside binding and recognition, and shed light on its putative galactomannan-binding site. Coordinates and observed structure factor amplitudes have been deposited with the Protein Data Bank (4CVU and 4UOJ). The T. harzianum β-mannosidase 2A nucleotide sequence has GenBank accession number BankIt1712036 GeneMark.hmm KJ624918.
- Instituto de Física de São Carlos, Universidade de São Paulo, Brazil.
Organizational Affiliation: