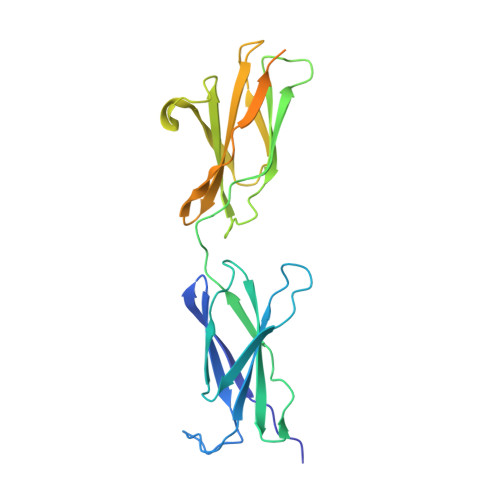
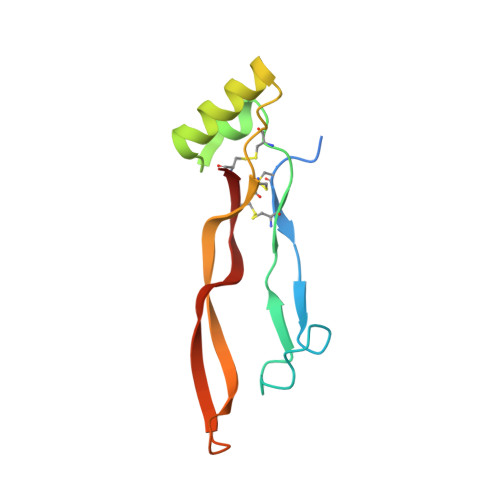
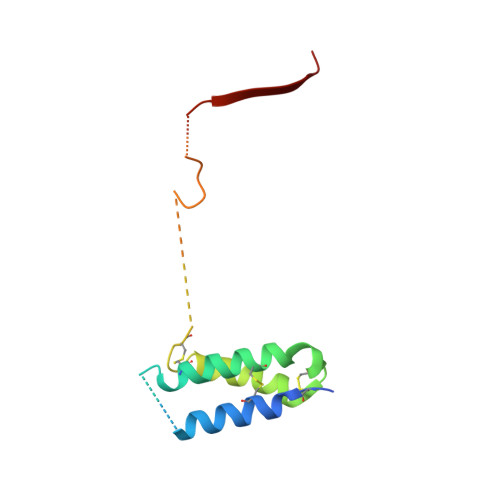
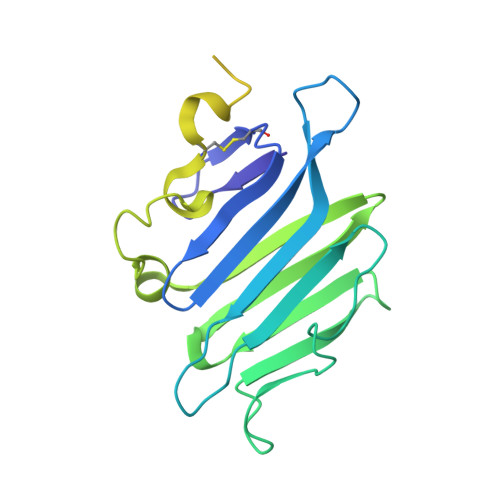
Repulsive Guidance Molecule is a Structural Bridge between Neogenin and Bone Morphogenetic Protein.
Healey, E.G., Bishop, B., Elegheert, J., Bell, C.H., Padilla-Parra, S., Siebold, C.(2015) Nat Struct Mol Biol 22: 458
- PubMed: 25938661 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nsmb.3016
- Primary Citation Related Structures:
4UHY, 4UHZ, 4UI0, 4UI1, 4UI2 - PubMed Abstract:
Repulsive guidance molecules (RGMs) control crucial processes including cell motility, adhesion, immune-cell regulation and systemic iron metabolism. RGMs signal via the neogenin (NEO1) and the bone morphogenetic protein (BMP) pathways. Here, we report crystal structures of the N-terminal domains of all human RGM family members in complex with the BMP ligand BMP2, revealing a new protein fold and a conserved BMP-binding mode. Our structural and functional data suggest a pH-linked mechanism for RGM-activated BMP signaling and offer a rationale for RGM mutations causing juvenile hemochromatosis. We also determined the crystal structure of the ternary BMP2-RGM-NEO1 complex, which, along with solution scattering and live-cell super-resolution fluorescence microscopy, indicates BMP-induced clustering of the RGM-NEO1 complex. Our results show how RGM acts as the central hub that links BMP and NEO1 and physically connects these fundamental signaling pathways.
- Division of Structural Biology, Wellcome Trust Centre for Human Genetics, University of Oxford, Oxford, UK.
Organizational Affiliation: