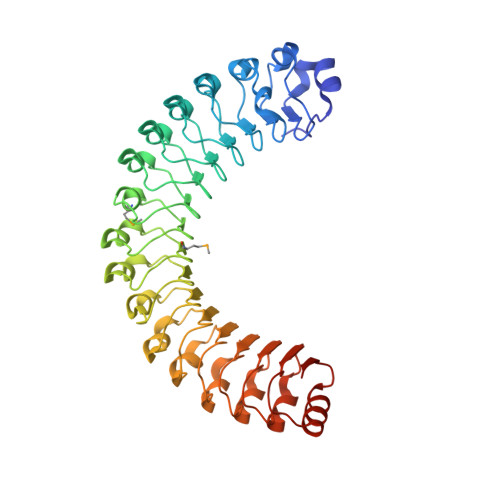
Structural characterization of a novel subfamily of leucine-rich repeat proteins from the human pathogen Leptospira interrogans.
Miras, I., Saul, F., Nowakowski, M., Weber, P., Haouz, A., Shepard, W., Picardeau, M.(2015) Acta Crystallogr D Biol Crystallogr 71: 1351-1359
- PubMed: 26057675 Search on PubMed
- DOI: https://doi.org/10.1107/S139900471500704X
- Primary Citation Related Structures:
4TZH, 4U06, 4U08, 4U09 - PubMed Abstract:
Pathogenic Leptospira spp. are the agents of leptospirosis, an emerging zoonotic disease. Analyses of Leptospira genomes have shown that the pathogenic leptospires (but not the saprophytes) possess a large number of genes encoding proteins containing leucine-rich repeat (LRR) domains. In other pathogenic bacteria, proteins with LRR domains have been shown to be involved in mediating host-cell attachment and invasion, but their functions remain unknown in Leptospira. To gain insight into the potential function of leptospiral LRR proteins, the crystal structures of four LRR proteins that represent a novel subfamily with consecutive stretches of a 23-amino-acid LRR repeat motif have been solved. The four proteins analyzed adopt the characteristic α/β-solenoid horseshoe fold. The exposed residues of the inner concave surfaces of the solenoid, which constitute a putative functional binding site, are not conserved. The various leptospiral LRR proteins could therefore recognize distinct structural motifs of different host proteins and thus serve separate and complementary functions in the physiology of these bacteria.
- Institut Pasteur, Plate-forme de Cristallographie, CNRS-UMR 3528, Paris, France.
Organizational Affiliation: