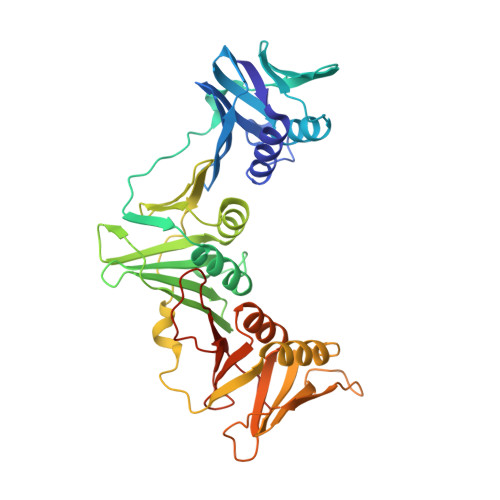
Structural insight into beta-Clamp and its interaction with DNA Ligase in Helicobacter pylori.
Pandey, P., Tarique, K.F., Mazumder, M., Rehman, S.A., Kumari, N., Gourinath, S.(2016) Sci Rep 6: 31181-31181
- PubMed: 27499105 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/srep31181
- Primary Citation Related Structures:
4RKI, 4S3I, 5FRQ - PubMed Abstract:
Helicobacter pylori, a gram-negative and microaerophilic bacterium, is the major cause of chronic gastritis, gastric ulcers and gastric cancer. Owing to its central role, DNA replication machinery has emerged as a prime target for the development of antimicrobial drugs. Here, we report 2Å structure of β-clamp from H. pylori (Hpβ-clamp), which is one of the critical components of DNA polymerase III. Despite of similarity in the overall fold of eubacterial β-clamp structures, some distinct features in DNA interacting loops exists that have not been reported previously. The in silico prediction identified the potential binders of β-clamp such as alpha subunit of DNA pol III and DNA ligase with identification of β-clamp binding regions in them and validated by SPR studies. Hpβ-clamp interacts with DNA ligase in micromolar binding affinity. Moreover, we have successfully determined the co-crystal structure of β-clamp with peptide from DNA ligase (not reported earlier in prokaryotes) revealing the region from ligase that interacts with β-clamp.
- School of Life Sciences, Jawaharlal Nehru University, New Delhi, India.
Organizational Affiliation: