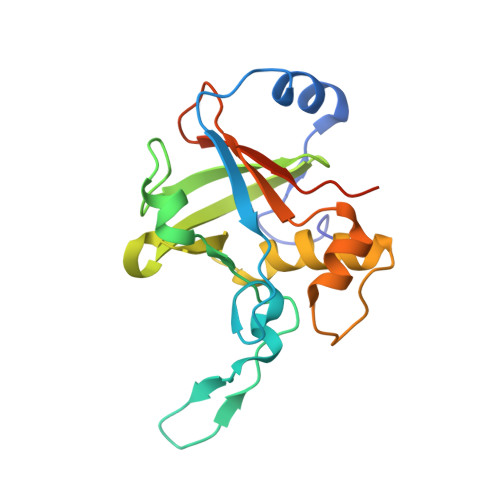
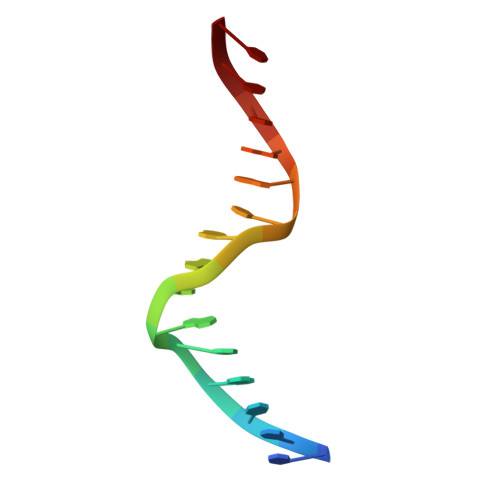
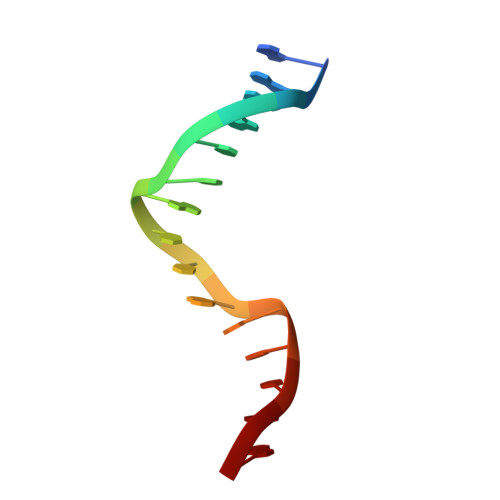
Crystal structure of the R-protein of the multisubunit ATP-dependent restriction endonuclease NgoAVII.
Tamulaitiene, G., Silanskas, A., Grazulis, S., Zaremba, M., Siksnys, V.(2014) Nucleic Acids Res 42: 14022-14030
- PubMed: 25429979 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gku1237
- Primary Citation Related Structures:
4RCT, 4RD5, 4RDM - PubMed Abstract:
The restriction endonuclease (REase) NgoAVII is composed of two proteins, R.NgoAVII and N.NgoAVII, and shares features of both Type II restriction enzymes and Type I/III ATP-dependent restriction enzymes (see accompanying paper Zaremba et al., 2014). Here we present crystal structures of the R.NgoAVII apo-protein and the R.NgoAVII C-terminal domain bound to a specific DNA. R.NgoAVII is composed of two domains: an N-terminal nucleolytic PLD domain; and a C-terminal B3-like DNA-binding domain identified previously in BfiI and EcoRII REases, and in plant transcription factors. Structural comparison of the B3-like domains of R.NgoAVII, EcoRII, BfiI and the plant transcription factors revealed a conserved DNA-binding surface comprised of N- and C-arms that together grip the DNA. The C-arms of R.NgoAVII, EcoRII, BfiI and plant B3 domains are similar in size, but the R.NgoAVII N-arm which makes the majority of the contacts to the target site is much longer. The overall structures of R.NgoAVII and BfiI are similar; however, whilst BfiI has stand-alone catalytic activity, R.NgoAVII requires an auxiliary cognate N.NgoAVII protein and ATP hydrolysis in order to cleave DNA at the target site. The structures we present will help formulate future experiments to explore the molecular mechanisms of intersubunit crosstalk that control DNA cleavage by R.NgoAVII and related endonucleases.
- Department of Protein-DNA Interactions, Institute of Biotechnology, Vilnius University, Graiciuno 8, LT-02241 Vilnius, Lithuania.
Organizational Affiliation: