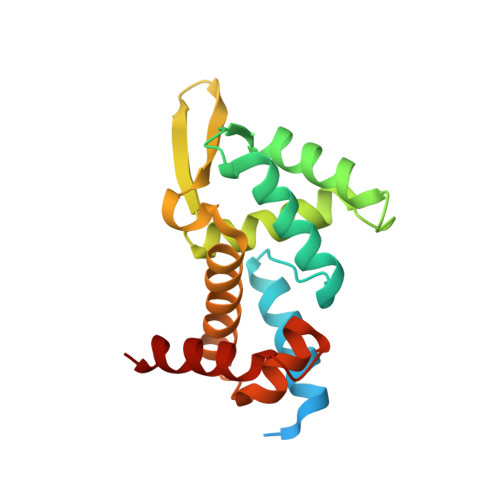
Structural basis for RNA recognition in roquin-mediated post-transcriptional gene regulation.
Schlundt, A., Heinz, G.A., Janowski, R., Geerlof, A., Stehle, R., Heissmeyer, V., Niessing, D., Sattler, M.(2014) Nat Struct Mol Biol 21: 671-678
- PubMed: 25026077 Search on PubMed
- DOI: https://doi.org/10.1038/nsmb.2855
- Primary Citation Related Structures:
4QI0, 4QI2 - PubMed Abstract:
Roquin function in T cells is essential for the prevention of autoimmune disease. Roquin interacts with the 3' untranslated regions (UTRs) of co-stimulatory receptors and controls T-cell activation and differentiation. Here we show that the N-terminal ROQ domain from mouse roquin adopts an extended winged-helix (WH) fold, which is sufficient for binding to the constitutive decay element (CDE) in the Tnf 3' UTR. The crystal structure of the ROQ domain in complex with a prototypical CDE RNA stem-loop reveals tight recognition of the RNA stem and its triloop. Surprisingly, roquin uses mainly non-sequence-specific contacts to the RNA, thus suggesting a relaxed CDE consensus and implicating a broader spectrum of target mRNAs than previously anticipated. Consistently with this, NMR and binding experiments with CDE-like stem-loops together with cell-based assays confirm roquin-dependent regulation of relaxed CDE consensus motifs in natural 3' UTRs.
- 1] Biomolecular NMR Spectroscopy, Institute of Structural Biology, Helmholtz Zentrum München, Neuherberg, Germany. [2] Center for Integrated Protein Science Munich at Biomolecular NMR Spectroscopy, Department Chemie, Technische Universität München, Garching, Germany. [3].
Organizational Affiliation: