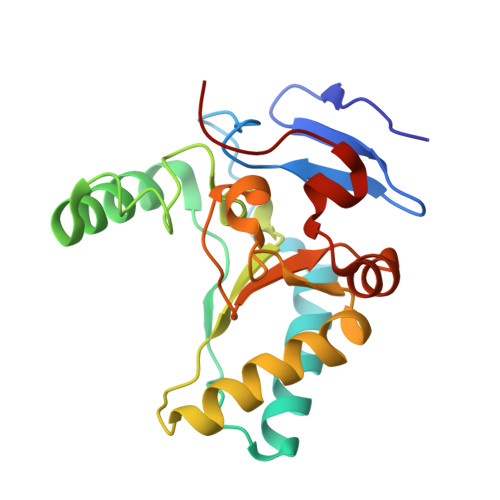
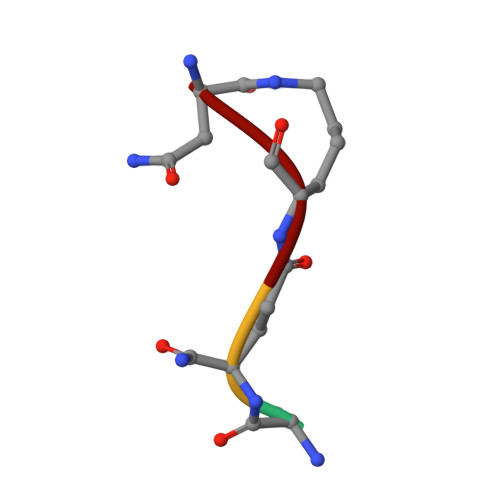
Structure of the LdcB LD-carboxypeptidase reveals the molecular basis of peptidoglycan recognition.
Hoyland, C.N., Aldridge, C., Cleverley, R.M., Duchene, M.C., Minasov, G., Onopriyenko, O., Sidiq, K., Stogios, P.J., Anderson, W.F., Daniel, R.A., Savchenko, A., Vollmer, W., Lewis, R.J.(2014) Structure 22: 949-960
- PubMed: 24909784 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2014.04.015
- Primary Citation Related Structures:
4JID, 4MPH, 4OX3, 4OX5, 4OXD - PubMed Abstract:
Peptidoglycan surrounds the bacterial cytoplasmic membrane to protect the cell against osmolysis. The biosynthesis of peptidoglycan, made of glycan strands crosslinked by short peptides, is the target of antibiotics like β-lactams and glycopeptides. Nascent peptidoglycan contains pentapeptides that are trimmed by carboxypeptidases to tetra- and tripeptides. The well-characterized DD-carboxypeptidases hydrolyze the terminal D-alanine from the stem pentapeptide to produce a tetrapeptide. However, few LD-carboxypeptidases that produce tripeptides have been identified, and nothing is known about substrate specificity in these enzymes. We report biochemical properties and crystal structures of the LD-carboxypeptidases LdcB from Streptococcus pneumoniae, Bacillus anthracis, and Bacillus subtilis. The enzymes are active against bacterial cell wall tetrapeptides and adopt a zinc-carboxypeptidase fold characteristic of the LAS superfamily. We have also solved the structure of S. pneumoniae LdcB with a product mimic, elucidating the residues essential for peptidoglycan recognition and the conformational changes that occur on ligand binding.
- Institute for Cell and Molecular Biosciences, Newcastle University, Newcastle upon Tyne NE2 4HH, UK.
Organizational Affiliation: