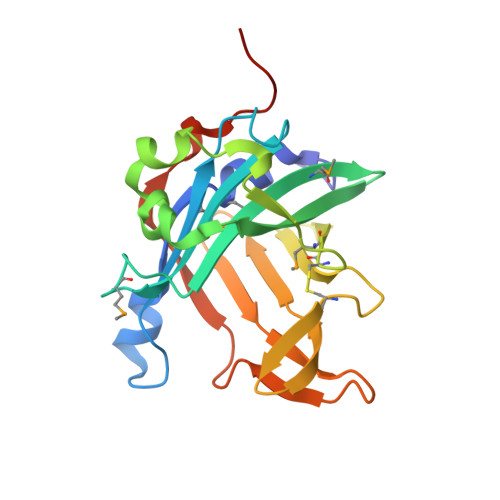
Structure and Mechanism of the Phycobiliprotein Lyase CpcT.
Zhou, W., Ding, W.L., Zeng, X.L., Dong, L.L., Zhao, B., Zhou, M., Scheer, H., Zhao, K.H., Yang, X.(2014) J Biological Chem 289: 26677-26689
- PubMed: 25074932 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M114.586743
- Primary Citation Related Structures:
4O4O, 4O4S - PubMed Abstract:
Pigmentation of light-harvesting phycobiliproteins of cyanobacteria requires covalent attachment of open-chain tetrapyrroles, bilins, to the apoproteins. Thioether formation via addition of a cysteine residue to the 3-ethylidene substituent of bilins is mediated by lyases. T-type lyases are responsible for attachment to Cys-155 of phycobiliprotein β-subunits. We present crystal structures of CpcT (All5339) from Nostoc (Anabaena) sp. PCC 7120 and its complex with phycocyanobilin at 1.95 and 2.50 Å resolution, respectively. CpcT forms a dimer and adopts a calyx-shaped β-barrel fold. Although the overall structure of CpcT is largely retained upon chromophore binding, arginine residues at the opening of the binding pocket undergo major rotameric rearrangements anchoring the propionate groups of phycocyanobilin. Based on the structure and mutational analysis, a reaction mechanism is proposed that accounts for chromophore stabilization and regio- and stereospecificity of the addition reaction. At the dimer interface, a loop extending from one subunit partially shields the opening of the phycocyanobilin binding pocket in the other subunit. Deletion of the loop or disruptions of the dimer interface significantly reduce CpcT lyase activity, suggesting functional relevance of the dimer. Dimerization is further enhanced by chromophore binding. The chromophore is largely buried in the dimer, but in the monomer, the 3-ethylidene group is accessible for the apophycobiliprotein, preferentially from the chromophore α-side. Asp-163 and Tyr-65 at the β- and α-face near the E-configured ethylidene group, respectively, support the acid-catalyzed nucleophilic Michael addition of cysteine 155 of the apoprotein to an N-acylimmonium intermediate proposed by Grubmayr and Wagner (Grubmayr, K., and Wagner, U. G. (1988) Monatsh. Chem. 119, 965-983).
- State Key Laboratory of Agricultural Microbiology, Huazhong Agricultural University, Wuhan 430070, China.
Organizational Affiliation: