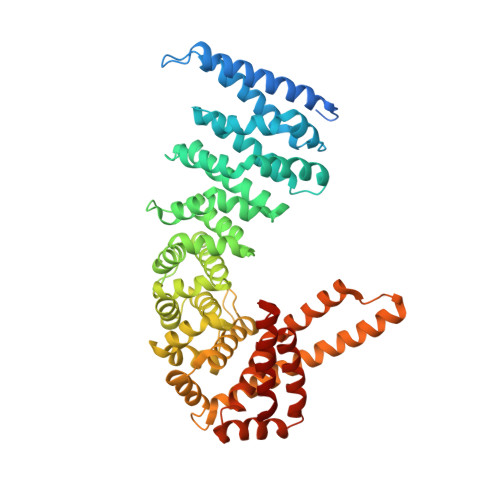
Structural insights into the novel ARM-repeat protein CTNNBL1 and its association with the hPrp19-CDC5L complex
Ahn, J.W., Kim, S., Kim, E.J., Kim, Y.J., Kim, K.J.(2014) Acta Crystallogr D Biol Crystallogr 70: 780-788
- PubMed: 24598747 Search on PubMed
- DOI: https://doi.org/10.1107/S139900471303318X
- Primary Citation Related Structures:
4MFU, 4MFV - PubMed Abstract:
The hPrp19-CDC5L complex plays a crucial role during human pre-mRNA splicing by catalytic activation of the spliceosome. In order to elucidate the molecular architecture of the hPrp19-CDC5L complex, the crystal structure of CTNNBL1, one of the major components of this complex, was determined. Unlike canonical ARM-repeat proteins such as β-catenin and importin-α, CTNNBL1 was found to contain a twisted and extended ARM-repeat structure at the C-terminal domain and, more importantly, the protein formed a stable dimer. A highly negatively charged patch formed in the N-terminal ARM-repeat domain of CTNNBL1 provides a binding site for CDC5L, a binding partner of the protein in the hPrp19-CDC5L complex, and these two proteins form a complex with a stoichiometry of 2:2. These findings not only present the crystal structure of a novel ARM-repeat protein, CTNNBL1, but also provide insights into the detailed molecular architecture of the hPrp19-CDC5L complex.
- Structural and Molecular Biology Laboratory, School of Life Sciences and Biotechnology, Kyungpook National University, Daehak-ro 80, Buk-ku, Daegu 702-701, Republic of Korea.
Organizational Affiliation: