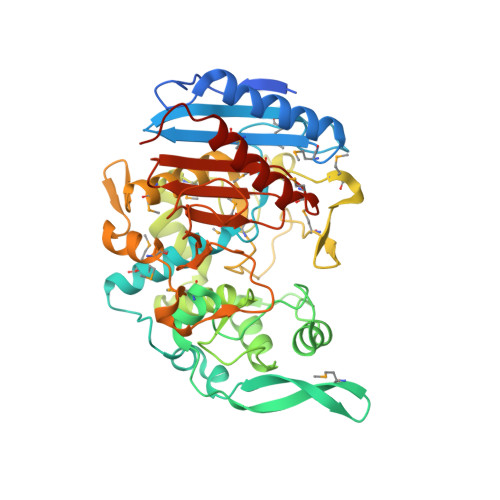
Structural basis for the beta-lactamase activity of EstU1, a family VIII carboxylesterase.
Cha, S.S., An, Y.J., Jeong, C.S., Kim, M.K., Jeon, J.H., Lee, C.M., Lee, H.S., Kang, S.G., Lee, J.H.(2013) Proteins 81: 2045-2051
- PubMed: 23737193 Search on PubMed
- DOI: https://doi.org/10.1002/prot.24334
- Primary Citation Related Structures:
4IVI, 4IVK - PubMed Abstract:
EstU1 is a unique family VIII carboxylesterase that displays hydrolytic activity toward the amide bond of clinically used β-lactam antibiotics as well as the ester bond of p-nitrophenyl esters. EstU1 assumes a β-lactamase-like modular architecture and contains the residues Ser100, Lys103, and Tyr218, which correspond to the three catalytic residues (Ser64, Lys67, and Tyr150, respectively) of class C β-lactamases. The structure of the EstU1/cephalothin complex demonstrates that the active site of EstU1 is not ideally tailored to perform an efficient deacylation reaction during the hydrolysis of β-lactam antibiotics. This result explains the weak β-lactamase activity of EstU1 compared with class C β-lactamases. Finally, structural and sequential comparison of EstU1 with other family VIII carboxylesterases elucidates an operative molecular strategy used by family VIII carboxylesterases to extend their substrate spectrum.
- Marine Biotechnology Research Division, Korea Institute of Ocean Science and Technology, Ansan, 426-744, Republic of Korea; Ocean Science and Technology School, Korea Maritime University, Pusan, 606-791, Republic of Korea; Department of Marine Biotechnology, University of Science and Technology, Daejeon, 305-333, Republic of Korea.
Organizational Affiliation: