TBA
Chi, B., Zaccai, N.R., Brady, R.L., Woolfson, D.N.(null) J Am Chem Soc
Experimental Data Snapshot
Starting Model: experimental
View more details
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| CC-Hex-II-Phi22 | 33 | synthetic construct | Mutation(s): 0 | 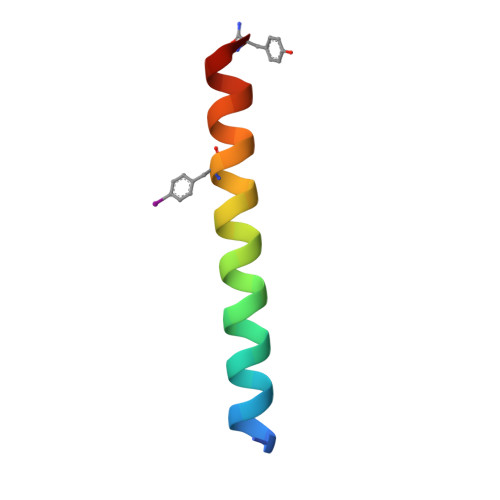 | |
| Ligands 2 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| GOL Download:Ideal Coordinates CCD File | D [auth A] | GLYCEROL C3 H8 O3 PEDCQBHIVMGVHV-UHFFFAOYSA-N |  | ||
| TBU Download:Ideal Coordinates CCD File | C [auth A], E [auth A], F [auth B] | TERTIARY-BUTYL ALCOHOL C4 H10 O DKGAVHZHDRPRBM-UHFFFAOYSA-N |  | ||
| Modified Residues 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Type | Formula | 2D Diagram | Parent |
| PHI Query on PHI | A, B | L-PEPTIDE LINKING | C9 H10 I N O2 |  | PHE |
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 46.67 | α = 90 |
| b = 46.67 | β = 90 |
| c = 94.06 | γ = 90 |
| Software Name | Purpose |
|---|---|
| REFMAC | refinement |
| PHENIX | refinement |