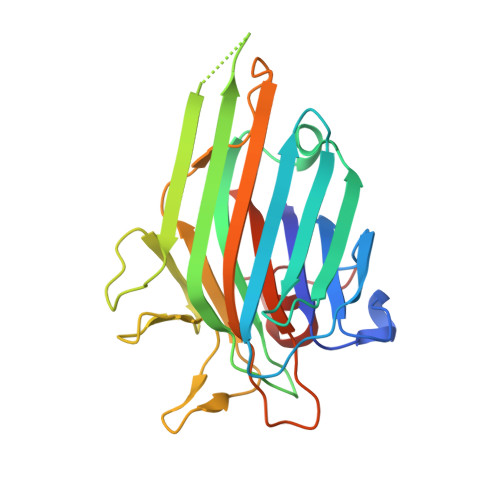
Crystal structure of Canavalia brasiliensis seed lectin (ConBr) in complex with beta-d-ribofuranose
Salviano, E., Rocha, B.A.M., Cavada, B.S., Delatorre, P., Santi-Gadelha, T., Gadelha, C.A.A., Silva-Filho, J.C., Nobrega, R.B., Farias, D.L.To be published.