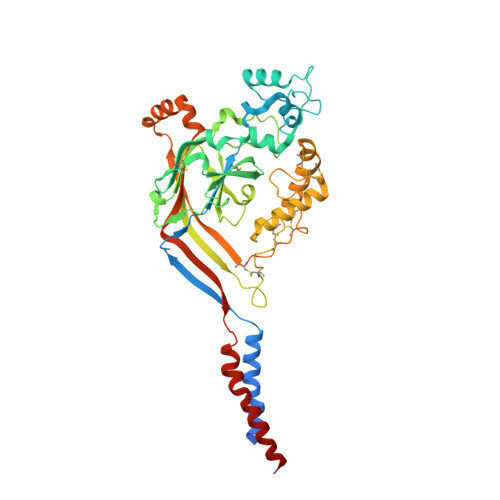
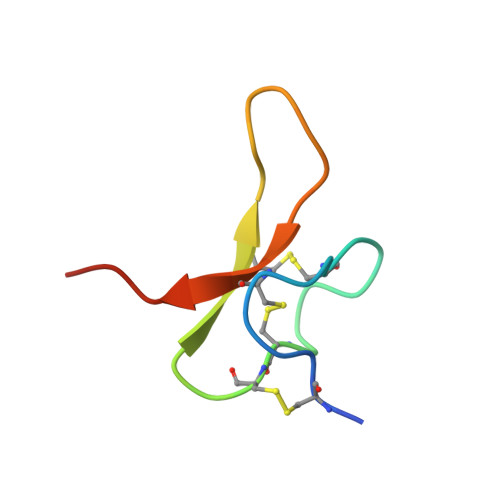
Structural plasticity and dynamic selectivity of acid-sensing ion channel-spider toxin complexes.
Baconguis, I., Gouaux, E.(2012) Nature 489: 400-405
- PubMed: 22842900 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nature11375
- Primary Citation Related Structures:
4FZ0, 4FZ1 - PubMed Abstract:
Acid-sensing ion channels (ASICs) are voltage-independent, amiloride-sensitive channels involved in diverse physiological processes ranging from nociception to taste. Despite the importance of ASICs in physiology, we know little about the mechanism of channel activation. Here we show that psalmotoxin activates non-selective and Na(+)-selective currents in chicken ASIC1a at pH 7.25 and 5.5, respectively. Crystal structures of ASIC1a-psalmotoxin complexes map the toxin binding site to the extracellular domain and show how toxin binding triggers an expansion of the extracellular vestibule and stabilization of the open channel pore. At pH 7.25 the pore is approximately 10 Å in diameter, whereas at pH 5.5 the pore is largely hydrophobic and elliptical in cross-section with dimensions of approximately 5 by 7 Å, consistent with a barrier mechanism for ion selectivity. These studies define mechanisms for activation of ASICs, illuminate the basis for dynamic ion selectivity and provide the blueprints for new therapeutic agents.
- Vollum Institute, Oregon Health and Science University, 3181 SW Sam Jackson Park Road, Portland, Oregon 97239, USA.
Organizational Affiliation: