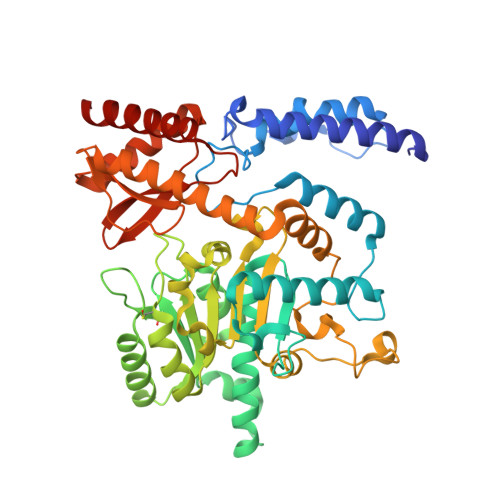
Structural study reveals that Ser-354 determines substrate specificity on human histidine decarboxylase
Komori, H., Nitta, Y., Ueno, H., Higuchi, Y.(2012) J Biological Chem 287: 29175-29183
- PubMed: 22767596 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M112.381897
- Primary Citation Related Structures:
4E1O - PubMed Abstract:
Histamine is an important chemical mediator for a wide variety of physiological reactions. L-histidine decarboxylase (HDC) is the primary enzyme responsible for histamine synthesis and produces histamine from histidine in a one-step reaction. In this study, we determined the crystal structure of human HDC (hHDC) complexed with the inhibitor histidine methyl ester. This structure shows the detailed features of the pyridoxal-5'-phosphate inhibitor adduct (external aldimine) at the active site of HDC. Moreover, a comparison of the structures of hHDC and aromatic L-amino acid (L-DOPA) decarboxylase showed that Ser-354 was a key residue for substrate specificity. The S354G mutation at the active site enlarged the size of the hHDC substrate-binding pocket and resulted in a decreased affinity for histidine, but an acquired ability to bind and act on L-DOPA as a substrate. These data provide insight into the molecular basis of substrate recognition among the group II pyridoxal-5'-phosphate-dependent decarboxylases.
- Department of Life Science, Graduate School of Life Science, University of Hyogo, 3-2-1 Koto, Kamigori-cho, Ako-gun, Hyogo 678-1297, Japan. komori@sci.u-hyogo.ac.jp
Organizational Affiliation: