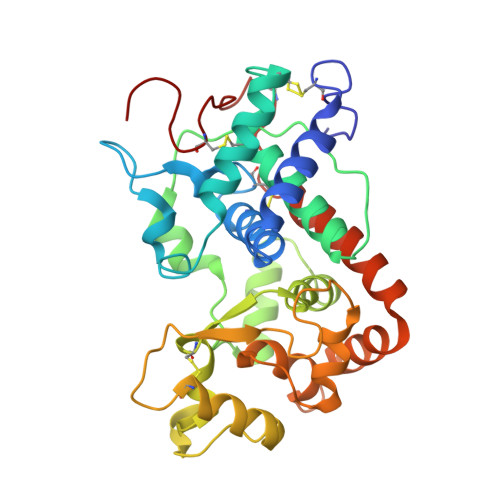
Post-Translational Modification and Extended Glycosylation Pattern of a Plant Latex Peroxidase of Native Source Characterized by X-Ray Crystallography.
Palm, G.J., Sharma, A., Kumari, M., Panjikar, S., Albrecht, D., Jagannadham, M.V., Hinrichs, W.(2014) FEBS J 281: 4319
- PubMed: 24980207 Search on PubMed
- DOI: https://doi.org/10.1111/febs.12900
- Primary Citation Related Structures:
4CUO - PubMed Abstract:
The crystal structure of banyan peroxidase purified from the latex of Ficus benghalensis has been solved at 1.67 Å resolution by single-wavelength anomalous diffraction phasing. The refined structure includes 306 amino acid residues, a heme and two calcium ions. The protein belongs to class III peroxidases and is the first one from plant latex. Extensive glycosylation was observed with N-linked glycans attached to seven asparagine residues. The enzyme is stable with respect to a wide pH range, temperature, chemical denaturants and organic solvents, probably as a result of its high glycosylation. An unexpected post-translational modification of Asp290 was identified as succinimide moiety. Kinetic parameters of banyan peroxidase have been determined using various hydrogen donor substrates and hydrogen peroxide. Coordinates and structure factors have been deposited in the Protein Data Bank under accession number 4CUO.
- Institut für Biochemie, Universität Greifswald, Germany.
Organizational Affiliation: